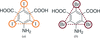issue contents
April 2010 issue
Experimental phasing and radiation damage
Proceedings of the CCP4 study weekend

Cover illustration: Speakers at the CCP4 Study Weekend 2009. Front row, left to right: James Holton, Arwen Pearson, Zbigniew Dauter, Elspeth Garman, Tobias Beck, Zbyszek Otwinowski. Middle row, left to right: Alexander Popov, Sean McSweeney, Garry Taylor, Helen Walden, Karthik Paithankar, Marc Schiltz, Gerard Bricogne. Back row, left to right: Kevin Cowtan, Martin Weik, Clemens Vonrhein, Peter Sun, Douglas Juers, George Sheldrick, Susan Lea, Airlie McCoy.
research papers
Open  access
access
 access
accessThis introductory paper to the CCP4 weekend on experimental phasing introduces the concept of the `phase problem' for non-experts. Modern methods of phasing are explored, including some recent examples that can be downloaded as tutorials.
Open  access
access
 access
accessThe basic causes of the radiation damage inflicted on macromolecular crystals during diffraction experiments are summarized, as well as the current state of research which attempts to understand and to mitigate it.
Open  access
access
 access
accessAn overview of techniques for recombinant incorporation of selenium and subsequent purification and crystallization of the resulting labelled protein.
Open  access
access
 access
accessIn order to overcome the difficulties associated with the `classical' heavy-atom derivatization procedure, an attempt has been made to develop a rational crystal-free heavy-atom-derivative screening method and a quick-soak derivatization procedure which allows heavy-atom compound identification.
Open  access
access
 access
accessMeasurements of the average thermal contractions (294→72 K) of 26 different cryosolutions are presented and discussed in conjunction with other recent advances in the rational design of protocols for cryogenic cooling in macromolecular crystallography.
Open  access
access
 access
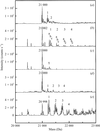
access5-Amino-2,4,6-tribromoisophthalic acid is used as a phasing tool for protein structure determination by MAD phasing. It is the second representative of a novel class of compounds for heavy-atom derivatization that combine heavy atoms with amino and carboxyl groups for binding to proteins.
Open  access
access
 access
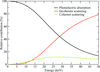
accessThe program RADDOSE computes the dose absorbed by a macromolecular crystal and here a guide is provided to help to ensure the proper use of the program. In the new version (v.3) described here, modifications to include the energy deposited owing to Compton scattering have been made.
Open  access
access
 access
accessDiffraction data collection parameters leading to optimal data quality are discussed in the context of different applications of these data.
Open  access
access
 access
accessA formula for absolute scattering power is derived to include spot fading arising from radiation damage and the crystal volume needed to collect diffraction data to a given resolution is calculated.
Open  access
access
 access
accessSoftware implementing a new method for the optimal choice of data-collection parameters, accounting for the effects of radiation damage, is presented.
Open  access
access
 access
accessCombining experimental phases and those from refinement of very incomplete models significantly improves electron-density maps.
Open  access
access
 access
accessRadiation-induced decay of crystal diffraction and additional specific chemical changes of macromolecules forming the crystal lattice are currently two of the main limiting factors in the acquisition of macromolecular diffraction data and macromolecular structure determination. Data-processing and phasing protocols are discussed in the context of radiation-induced changes.
Open  access
access
 access
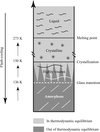
accessThe dynamical behaviour of crystalline macromolecules and their surrounding solvent as a function of cryo-temperature is reviewed.
Open  access
access
 access
accessSite-specific radiation damage and anisotropy of anomalous scattering can induce intensity differences in symmetry-related reflections. If the data are kept unmerged, these symmetry-breaking effects can become a source of phase information.
Open  access
access
 access
accessThe pitfalls of experimental phasing are described.
Open  access
access
 access
accessSeveral new methods are evaluated for use in the improvement of experimental phases in the framework of a classical density-modification calculation. These methods have been implemented in a new computer program, Parrot.
Open  access
access
 access
accessExperimental phasing with SHELXC/D/E has been enhanced by the incorporation of main-chain tracing into the iterative density modification; this also provides a simple and effective way of exploiting noncrystallographic symmetry.
Open  access
access
 access
accessCoot is a molecular-graphics program designed to assist in the building of protein and other macromolecular models. The current state of development and available features are presented.


 journal menu
journal menu