The centrosymmetric title compound, C
16H
10N
4O
2, is an organic red pigment utilized for H
2 gas sensors. The centrosymmetric diketopyrrolopyrrole moieties are connected by bifurcated N

H

O hydrogen bonds to form a ribbon structure along [110] and [1
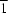
0] alternately. The molecules are stacked in a `hunter's fence' (
viz. when viewed from the side, molecules, slipped by 45° within molecular stacks, cross each other in a fence-like structure) fashion along the
a axis.
Supporting information
CCDC reference: 264100
Key indicators
- Single-crystal X-ray study
- T = 93 K
- Mean
![[sigma]](//journals.iucr.org/logos/entities/sigma_rmgif.gif) (C-C) = 0.010 Å
(C-C) = 0.010 Å
- R factor = 0.241
- wR factor = 0.565
- Data-to-parameter ratio = 10.9
checkCIF/PLATON results
No syntax errors found
 Alert level A
ABSTM02_ALERT_3_A The ratio of expected to reported Tmax/Tmin(RR') is < 0.50
Tmin and Tmax reported: 0.009 0.982
Tmin' and Tmax expected: 0.825 0.982
RR' = 0.011
Please check that your absorption correction is appropriate.
Alert level A
ABSTM02_ALERT_3_A The ratio of expected to reported Tmax/Tmin(RR') is < 0.50
Tmin and Tmax reported: 0.009 0.982
Tmin' and Tmax expected: 0.825 0.982
RR' = 0.011
Please check that your absorption correction is appropriate.
| Author Response: The single crystal used for measurements was extremely small and
thin (0.2x0.1x0.02 mm^3).
|
RFACG01_ALERT_3_A The value of the R factor is > 0.20
R factor given 0.241
| Author Response: The single crystal used for the analysis was extremely small and
and thin (0.2x0.1x0.02 mm^3). In addition, the crystal was slightly curved and
the crystallinity was rather poor. Nevertheless, this was the maximum which
we could do.
|
RFACR01_ALERT_3_A The value of the weighted R factor is > 0.45
Weighted R factor given 0.565
| Author Response: The single crystal used for the analysis was extremely small and
and thin (0.2x0.1x0.02 mm^3). In addition, the crystal was slightly curved and
the crystallinity was rather poor. Nevertheless, this was the maximum which
we could do.
|
RINTA01_ALERT_3_A The value of Rint is greater than 0.20
Rint given 0.246
| Author Response: The single crystal used for the analysis was extremely small and
and thin (0.2x0.1x0.02 mm^3). In addition, the crystal was slightly curved and
the crystallinity was rather poor. Nevertheless, this was the maximum which
we could do.
|
PLAT020_ALERT_3_A The value of Rint is greater than 0.10 ......... 0.25
| Author Response: The single crystal used for the analysis was extremely small and
and thin (0.2x0.1x0.02 mm^3). In addition, the crystal was slightly curved and
the crystallinity was rather poor. Nevertheless, this was the maximum which
we could do.
|
PLAT061_ALERT_3_A Tmax/Tmin Range Test RR' too Large ............. 0.01
| Author Response: The single crystal used for measurements was extremely small and
thin (0.2x0.1x0.02 mm^3).
|
PLAT082_ALERT_2_A High R1 Value .................................. 0.24
| Author Response: The single crystal used for the analysis was extremely small and
and thin (0.2x0.1x0.02 mm^3). In addition, the crystal was slightly curved and
the crystallinity was rather poor. Nevertheless, this was the maximum which
we could do.
|
PLAT084_ALERT_2_A High R2 Value .................................. 0.56
| Author Response: The single crystal used for the analysis was extremely small and
and thin (0.2x0.1x0.02 mm^3). In addition, the crystal was slightly curved and
the crystallinity was rather poor. Nevertheless, this was the maximum which
we could do.
|
 Alert level B
DIFMN02_ALERT_2_B The minimum difference density is < -0.1*ZMAX*1.00
_refine_diff_density_min given = -1.400
Test value = -0.800
DIFMX01_ALERT_2_B The maximum difference density is > 0.1*ZMAX*1.00
_refine_diff_density_max given = 1.600
Test value = 0.800
PLAT029_ALERT_3_B _diffrn_measured_fraction_theta_full Low ....... 0.96
PLAT097_ALERT_2_B Maximum (Positive) Residual Density ............ 1.60 e/A 3
PLAT098_ALERT_2_B Minimum (Negative) Residual Density ............ -1.40 e/A 3
Alert level B
DIFMN02_ALERT_2_B The minimum difference density is < -0.1*ZMAX*1.00
_refine_diff_density_min given = -1.400
Test value = -0.800
DIFMX01_ALERT_2_B The maximum difference density is > 0.1*ZMAX*1.00
_refine_diff_density_max given = 1.600
Test value = 0.800
PLAT029_ALERT_3_B _diffrn_measured_fraction_theta_full Low ....... 0.96
PLAT097_ALERT_2_B Maximum (Positive) Residual Density ............ 1.60 e/A 3
PLAT098_ALERT_2_B Minimum (Negative) Residual Density ............ -1.40 e/A 3
 Alert level C
DIFMN03_ALERT_1_C The minimum difference density is < -0.1*ZMAX*0.75
The relevant atom site should be identified.
DIFMX02_ALERT_1_C The minimum difference density is > 0.1*ZMAX*0.75
The relevant atom site should be identified.
RADNW01_ALERT_1_C The radiation wavelength lies outside the expected range
for the supplied radiation type. Expected range 1.54175-1.54180
Wavelength given = 1.54190
PLAT150_ALERT_1_C Volume as Calculated Differs from that Given ... 616.00 Ang-3
PLAT250_ALERT_2_C Large U3/U1 Ratio for Average U(i,j) Tensor .... 2.85
PLAT340_ALERT_3_C Low Bond Precision on C-C bonds (x 1000) Ang ... 10
Alert level C
DIFMN03_ALERT_1_C The minimum difference density is < -0.1*ZMAX*0.75
The relevant atom site should be identified.
DIFMX02_ALERT_1_C The minimum difference density is > 0.1*ZMAX*0.75
The relevant atom site should be identified.
RADNW01_ALERT_1_C The radiation wavelength lies outside the expected range
for the supplied radiation type. Expected range 1.54175-1.54180
Wavelength given = 1.54190
PLAT150_ALERT_1_C Volume as Calculated Differs from that Given ... 616.00 Ang-3
PLAT250_ALERT_2_C Large U3/U1 Ratio for Average U(i,j) Tensor .... 2.85
PLAT340_ALERT_3_C Low Bond Precision on C-C bonds (x 1000) Ang ... 10
8 ALERT level A = In general: serious problem
5 ALERT level B = Potentially serious problem
6 ALERT level C = Check and explain
0 ALERT level G = General alerts; check
4 ALERT type 1 CIF construction/syntax error, inconsistent or missing data
7 ALERT type 2 Indicator that the structure model may be wrong or deficient
8 ALERT type 3 Indicator that the structure quality may be low
0 ALERT type 4 Improvement, methodology, query or suggestion
Data collection: PROCESS-AUTO (Rigaku, 1998); cell refinement: PROCESS-AUTO; data reduction: TEXSAN (Molecular Structure Corporation, 2001); program(s) used to solve structure: SHELXS86 (Sheldrick, 1985); program(s) used to refine structure: TEXSAN; molecular graphics: ORTEPIII (Burnett & Johnson, 1996); software used to prepare material for publication: TEXSAN.
Crystal data top
| C16H10N4O2 | F(000) = 300.0 |
| Mr = 290.28 | Dx = 1.565 Mg m−3 |
| Monoclinic, P21/n | Cu Kα radiation, λ = 1.5419 Å |
| Hall symbol: -P 2yn | Cell parameters from 4967 reflections |
| a = 3.722 (1) Å | θ = 3.3–68.3° |
| b = 6.263 (3) Å | µ = 0.89 mm−1 |
| c = 26.506 (9) Å | T = 93 K |
| β = 94.41 (2)° | Platelet, red |
| V = 616.0 (4) Å3 | 0.20 × 0.10 × 0.02 mm |
| Z = 2 | |
Data collection top
Rigaku R-AXIS RAPID Imaging Plate
diffractometer | 711 reflections with F2 > 2σ(F2) |
| Detector resolution: 10.00 pixels mm-1 | Rint = 0.246 |
| 48 frames, δ ω = 15° scans | θmax = 68.2° |
Absorption correction: multi-scan
(ABSCOR; Higashi, 1995) | h = 0→4 |
| Tmin = 0.009, Tmax = 0.982 | k = 0→6 |
| 6710 measured reflections | l = −31→31 |
| 1089 independent reflections | |
Refinement top
| Refinement on F2 | H-atom parameters not refined |
| R[F2 > 2σ(F2)] = 0.241 | w = 1/[σ2(Fo2) + {0.187[Max(Fo2,0) + 2Fc2]/3}2] |
| wR(F2) = 0.565 | (Δ/σ)max < 0.001 |
| S = 1.78 | Δρmax = 1.60 e Å−3 |
| 1089 reflections | Δρmin = −1.40 e Å−3 |
| 100 parameters | |
Special details top
Refinement. Refinement using reflections with F2 > -3.0 σ(F2). The
weighted R-factor (wR) and goodness of fit (S) are based
on F2. R-factor (gt) are based on F. The threshold
expression of F2 > 2.0 σ(F2) is used only for calculating
R-factor (gt). |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2) top| | x | y | z | Uiso*/Ueq | |
| O1 | 0.858 (2) | 0.8404 (9) | 0.4440 (2) | 0.0509 | |
| N1 | 0.503 (2) | 0.707 (1) | 0.7167 (3) | 0.0541 | |
| N2 | 0.727 (2) | 0.7912 (10) | 0.5273 (2) | 0.0442 | |
| C1 | 0.541 (2) | 0.667 (1) | 0.6115 (3) | 0.0465 | |
| C2 | 0.647 (2) | 0.855 (1) | 0.6375 (3) | 0.0491 | |
| C3 | 0.630 (2) | 0.865 (1) | 0.6875 (3) | 0.0481 | |
| C4 | 0.398 (2) | 0.526 (1) | 0.6921 (3) | 0.0481 | |
| C5 | 0.405 (2) | 0.499 (1) | 0.6413 (3) | 0.0466 | |
| C6 | 0.560 (2) | 0.640 (1) | 0.5577 (3) | 0.0479 | |
| C7 | 0.455 (2) | 0.477 (1) | 0.5251 (3) | 0.0461 | |
| C8 | 0.730 (2) | 0.733 (1) | 0.4776 (3) | 0.0430 | |
| H1 | 0.8279 | 0.9212 | 0.5402 | 0.0531* | |
| H2 | 0.7307 | 0.9745 | 0.6195 | 0.0589* | |
| H3 | 0.7125 | 0.9926 | 0.7041 | 0.0577* | |
| H4 | 0.3136 | 0.4113 | 0.7114 | 0.0577* | |
| H5 | 0.3216 | 0.3699 | 0.6258 | 0.0559* | |
Atomic displacement parameters (Å2) top| | U11 | U22 | U33 | U12 | U13 | U23 |
| O1 | 0.0624 | 0.0262 | 0.0633 | −0.0000 | −0.0005 | −0.0004 |
| N1 | 0.0570 | 0.0282 | 0.0763 | −0.0013 | −0.0014 | −0.0015 |
| N2 | 0.0466 | 0.0181 | 0.0675 | −0.0012 | 0.0015 | −0.0050 |
| C1 | 0.0448 | 0.0193 | 0.0756 | 0.0026 | 0.0069 | 0.0028 |
| C2 | 0.0452 | 0.0260 | 0.0754 | 0.0051 | 0.0002 | 0.0036 |
| C3 | 0.0446 | 0.0292 | 0.0697 | 0.0029 | −0.0001 | 0.0082 |
| C4 | 0.0390 | 0.0310 | 0.0733 | 0.0071 | −0.0022 | −0.0007 |
| C5 | 0.0415 | 0.0256 | 0.0727 | −0.0031 | 0.0051 | −0.0010 |
| C6 | 0.0542 | 0.0284 | 0.0598 | −0.0044 | −0.0053 | 0.0009 |
| C7 | 0.0552 | 0.0136 | 0.0686 | 0.0028 | −0.0023 | 0.0001 |
| C8 | 0.0404 | 0.0274 | 0.0612 | 0.0013 | 0.0041 | 0.0016 |
Geometric parameters (Å, º) top
| O1—C8 | 1.240 (10) | C2—C3 | 1.33 (1) |
| N1—C3 | 1.36 (1) | C2—H2 | 0.950 |
| N1—C4 | 1.35 (1) | C3—H3 | 0.950 |
| N2—C6 | 1.42 (1) | C4—C5 | 1.36 (1) |
| N2—C8 | 1.367 (10) | C4—H4 | 0.950 |
| N2—H1 | 0.950 | C5—H5 | 0.950 |
| C1—C2 | 1.40 (1) | C6—C7 | 1.37 (1) |
| C1—C5 | 1.43 (1) | C7—C7i | 1.43 (2) |
| C1—C6 | 1.45 (1) | C7—C8i | 1.48 (1) |
| | | |
| O1···N2ii | 2.847 (8) | N2···C6iv | 3.28 (1) |
| O1···C7iii | 3.29 (1) | | |
| | | |
| C3—N1—C4 | 115.7 (7) | N1—C4—H4 | 118.1 |
| C6—N2—C8 | 113.8 (6) | C5—C4—H4 | 118.1 |
| C6—N2—H1 | 123.1 | C1—C5—C4 | 119.3 (8) |
| C8—N2—H1 | 123.1 | C1—C5—H5 | 120.4 |
| C2—C1—C5 | 116.4 (8) | C4—C5—H5 | 120.4 |
| C2—C1—C6 | 123.5 (8) | N2—C6—C1 | 122.4 (7) |
| C5—C1—C6 | 120.1 (7) | N2—C6—C7 | 104.8 (7) |
| C1—C2—C3 | 119.5 (8) | C1—C6—C7 | 132.7 (8) |
| C1—C2—H2 | 120.2 | C6—C7—C7i | 111.3 (9) |
| C3—C2—H2 | 120.2 | C6—C7—C8i | 142.8 (8) |
| N1—C3—C2 | 125.2 (8) | C7i—C7—C8i | 105.9 (8) |
| N1—C3—H3 | 117.4 | O1—C8—N2 | 125.4 (7) |
| C2—C3—H3 | 117.4 | O1—C8—C7i | 130.4 (7) |
| N1—C4—C5 | 123.8 (8) | N2—C8—C7i | 104.2 (6) |
| Symmetry codes: (i) −x+1, −y+1, −z+1; (ii) −x+2, −y+2, −z+1; (iii) −x+2, −y+1, −z+1; (iv) x+1, y, z. |
Hydrogen-bond geometry (Å, º) top
| D—H···A | D—H | H···A | D···A | D—H···A |
| N2—H1···O1ii | 0.95 | 1.92 | 2.847 (8) | 164 |
| Symmetry code: (ii) −x+2, −y+2, −z+1. |
 H
H O hydrogen bonds to form a ribbon structure along [110] and [1
O hydrogen bonds to form a ribbon structure along [110] and [1![[sigma]](//journals.iucr.org/logos/entities/sigma_rmgif.gif) (C-C) = 0.010 Å
(C-C) = 0.010 ÅAlert level A ABSTM02_ALERT_3_A The ratio of expected to reported Tmax/Tmin(RR') is < 0.50 Tmin and Tmax reported: 0.009 0.982 Tmin' and Tmax expected: 0.825 0.982 RR' = 0.011 Please check that your absorption correction is appropriate.
Alert level B DIFMN02_ALERT_2_B The minimum difference density is < -0.1*ZMAX*1.00 _refine_diff_density_min given = -1.400 Test value = -0.800 DIFMX01_ALERT_2_B The maximum difference density is > 0.1*ZMAX*1.00 _refine_diff_density_max given = 1.600 Test value = 0.800 PLAT029_ALERT_3_B _diffrn_measured_fraction_theta_full Low ....... 0.96 PLAT097_ALERT_2_B Maximum (Positive) Residual Density ............ 1.60 e/A 3 PLAT098_ALERT_2_B Minimum (Negative) Residual Density ............ -1.40 e/A 3
Alert level C DIFMN03_ALERT_1_C The minimum difference density is < -0.1*ZMAX*0.75 The relevant atom site should be identified. DIFMX02_ALERT_1_C The minimum difference density is > 0.1*ZMAX*0.75 The relevant atom site should be identified. RADNW01_ALERT_1_C The radiation wavelength lies outside the expected range for the supplied radiation type. Expected range 1.54175-1.54180 Wavelength given = 1.54190 PLAT150_ALERT_1_C Volume as Calculated Differs from that Given ... 616.00 Ang-3 PLAT250_ALERT_2_C Large U3/U1 Ratio for Average U(i,j) Tensor .... 2.85 PLAT340_ALERT_3_C Low Bond Precision on C-C bonds (x 1000) Ang ... 10

 journal menu
journal menu










 Alert level A
Alert level A
 Alert level B
Alert level B
 Alert level C
Alert level C



