The crystal structure of the title compound, [Cd(C2H3N4S)2]n, consists of octahedrally coordinated Cd atoms that are linked by the anionic ligands into a layer structure. The Cd atom lies on a special position of site symmetry 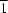 .
.
Supporting information
Crystallographic Information File (CIF) https://doi.org/10.1107/S160053680604699X/xu2163sup1.cif | |
Structure factor file (CIF format) https://doi.org/10.1107/S160053680604699X/xu2163Isup2.hkl |
CCDC reference: 630529
Key indicators
- Single-crystal X-ray study
- T = 295 K
- Mean
![[sigma]](//journals.iucr.org/logos/entities/sigma_rmgif.gif) (N-C) = 0.005 Å
(N-C) = 0.005 Å - R factor = 0.029
- wR factor = 0.078
- Data-to-parameter ratio = 15.2
checkCIF/PLATON results
No syntax errors found
Alert level B ABSTM02_ALERT_3_B The ratio of expected to reported Tmax/Tmin(RR') is < 0.75 Tmin and Tmax reported: 0.353 0.738 Tmin(prime) and Tmax expected: 0.643 0.724 RR(prime) = 0.539 Please check that your absorption correction is appropriate. PLAT061_ALERT_3_B Tmax/Tmin Range Test RR' too Large ............. 0.53
Alert level C PLAT062_ALERT_4_C Rescale T(min) & T(max) by ..................... 0.98 PLAT764_ALERT_4_C Overcomplete CIF Bond List Detected (Rep/Expd) . 1.15 Ratio
0 ALERT level A = In general: serious problem 2 ALERT level B = Potentially serious problem 2 ALERT level C = Check and explain 0 ALERT level G = General alerts; check 0 ALERT type 1 CIF construction/syntax error, inconsistent or missing data 0 ALERT type 2 Indicator that the structure model may be wrong or deficient 2 ALERT type 3 Indicator that the structure quality may be low 2 ALERT type 4 Improvement, methodology, query or suggestion 0 ALERT type 5 Informative message, check
Computing details top
Data collection: RAPID-AUTO (Rigaku, 2004); cell refinement: RAPID-AUTO; data reduction: CrystalStructure (Rigaku/MSC, 2004); program(s) used to solve structure: SHELXS97 (Sheldrick, 1997); program(s) used to refine structure: SHELXL97 (Sheldrick, 1997); molecular graphics: X-SEED (Barbour, 2001); software used to prepare material for publication: publCIF (Westrip, 2006).
Poly[bis(µ-1-methyl-1,2,3,4-tetrazol-5-ylthiolato)cadmium(II)] top
Crystal data top
| [Cd(C2H3N4S)2] | F(000) = 332 |
| Mr = 342.69 | Dx = 2.376 Mg m−3 |
| Monoclinic, P21/c | Mo Kα radiation, λ = 0.71073 Å |
| Hall symbol: -P 2ybc | Cell parameters from 4122 reflections |
| a = 11.431 (2) Å | θ = 3.1–27.5° |
| b = 5.395 (1) Å | µ = 2.69 mm−1 |
| c = 7.771 (2) Å | T = 295 K |
| β = 92.00 (3)° | Block, colorless |
| V = 478.95 (17) Å3 | 0.16 × 0.14 × 0.12 mm |
| Z = 2 |
Data collection top
| Rigaku R-AXIS RAPID IP diffractometer | 1082 independent reflections |
| Radiation source: fine-focus sealed tube | 991 reflections with I > 2σ(I) |
| Graphite monochromator | Rint = 0.059 |
| ω scans | θmax = 27.5°, θmin = 3.1° |
| Absorption correction: multi-scan (ABSCOR; Higashi, 1995) | h = −14→14 |
| Tmin = 0.353, Tmax = 0.738 | k = −7→6 |
| 4300 measured reflections | l = −9→10 |
Refinement top
| Refinement on F2 | Primary atom site location: structure-invariant direct methods |
| Least-squares matrix: full | Secondary atom site location: difference Fourier map |
| R[F2 > 2σ(F2)] = 0.029 | Hydrogen site location: inferred from neighbouring sites |
| wR(F2) = 0.078 | H-atom parameters constrained |
| S = 1.09 | w = 1/[σ2(Fo2) + (0.0399P)2 + 0.0392P] where P = (Fo2 + 2Fc2)/3 |
| 1082 reflections | (Δ/σ)max = 0.001 |
| 71 parameters | Δρmax = 0.60 e Å−3 |
| 0 restraints | Δρmin = −0.92 e Å−3 |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2) top
| x | y | z | Uiso*/Ueq | ||
| Cd1 | 0.5000 | 0.5000 | 0.5000 | 0.01851 (15) | |
| S1 | 0.60509 (5) | 0.71020 (11) | 0.23686 (8) | 0.01948 (18) | |
| N1 | 0.66177 (18) | 0.2888 (4) | 0.0520 (3) | 0.0200 (5) | |
| N2 | 0.7629 (2) | 0.1886 (4) | −0.0045 (3) | 0.0253 (5) | |
| N3 | 0.85182 (19) | 0.3246 (4) | 0.0416 (3) | 0.0255 (5) | |
| N4 | 0.8086 (3) | 0.5168 (3) | 0.1288 (4) | 0.0182 (6) | |
| C1 | 0.6914 (3) | 0.4953 (3) | 0.1363 (4) | 0.0160 (6) | |
| C2 | 0.8868 (2) | 0.6955 (5) | 0.2135 (4) | 0.0287 (6) | |
| H2A | 0.8470 | 0.8510 | 0.2246 | 0.043* | |
| H2B | 0.9548 | 0.7181 | 0.1461 | 0.043* | |
| H2C | 0.9103 | 0.6351 | 0.3257 | 0.043* |
Atomic displacement parameters (Å2) top
| U11 | U22 | U33 | U12 | U13 | U23 | |
| Cd1 | 0.0183 (2) | 0.0209 (2) | 0.0165 (2) | −0.00060 (7) | 0.00263 (13) | 0.00034 (7) |
| S1 | 0.0230 (3) | 0.0200 (4) | 0.0158 (3) | 0.0054 (2) | 0.0068 (2) | 0.0039 (2) |
| N1 | 0.0181 (11) | 0.0210 (11) | 0.0209 (12) | −0.0003 (8) | 0.0012 (8) | −0.0018 (8) |
| N2 | 0.0231 (12) | 0.0266 (12) | 0.0266 (13) | 0.0011 (9) | 0.0062 (9) | −0.0075 (9) |
| N3 | 0.0214 (12) | 0.0276 (12) | 0.0280 (13) | 0.0023 (8) | 0.0067 (10) | −0.0061 (9) |
| N4 | 0.0162 (13) | 0.0225 (14) | 0.0159 (14) | −0.0012 (7) | 0.0031 (10) | −0.0013 (7) |
| C1 | 0.0159 (15) | 0.0194 (17) | 0.0129 (17) | −0.0006 (7) | 0.0029 (12) | 0.0040 (7) |
| C2 | 0.0225 (14) | 0.0304 (16) | 0.0330 (17) | −0.0081 (10) | 0.0008 (12) | −0.0050 (11) |
Geometric parameters (Å, º) top
| Cd1—N1i | 2.441 (2) | N1—N2 | 1.363 (3) |
| Cd1—N1ii | 2.441 (2) | N1—Cd1iii | 2.441 (2) |
| Cd1—S1 | 2.6612 (9) | N2—N3 | 1.294 (3) |
| Cd1—S1iii | 2.6685 (8) | N3—N4 | 1.342 (3) |
| Cd1—S1iv | 2.6612 (9) | N4—C1 | 1.349 (4) |
| Cd1—S1v | 2.6685 (8) | N4—C2 | 1.456 (3) |
| S1—C1 | 1.727 (3) | C2—H2A | 0.9600 |
| S1—Cd1i | 2.6685 (8) | C2—H2B | 0.9600 |
| N1—C1 | 1.330 (3) | C2—H2C | 0.9600 |
| N1ii—Cd1—N1i | 180.0 | C1—N1—N2 | 106.7 (2) |
| N1ii—Cd1—S1iv | 87.73 (5) | C1—N1—Cd1iii | 142.6 (2) |
| N1i—Cd1—S1iv | 92.27 (5) | N2—N1—Cd1iii | 109.85 (15) |
| N1ii—Cd1—S1 | 92.27 (5) | N3—N2—N1 | 110.6 (2) |
| N1i—Cd1—S1 | 87.73 (5) | N2—N3—N4 | 106.2 (2) |
| S1iv—Cd1—S1 | 180.0 | N3—N4—C1 | 109.8 (2) |
| N1ii—Cd1—S1iii | 93.46 (5) | N3—N4—C2 | 120.6 (3) |
| N1i—Cd1—S1iii | 86.54 (5) | C1—N4—C2 | 129.3 (3) |
| S1iv—Cd1—S1iii | 94.35 (2) | N1—C1—N4 | 106.7 (2) |
| S1—Cd1—S1iii | 85.65 (2) | N1—C1—S1 | 130.2 (2) |
| N1ii—Cd1—S1v | 86.54 (5) | N4—C1—S1 | 123.10 (19) |
| N1i—Cd1—S1v | 93.46 (5) | N4—C2—H2A | 109.5 |
| S1iv—Cd1—S1v | 85.65 (2) | N4—C2—H2B | 109.5 |
| S1—Cd1—S1v | 94.35 (2) | H2A—C2—H2B | 109.5 |
| S1iii—Cd1—S1v | 180.0 | N4—C2—H2C | 109.5 |
| C1—S1—Cd1 | 110.00 (9) | H2A—C2—H2C | 109.5 |
| C1—S1—Cd1i | 109.37 (11) | H2B—C2—H2C | 109.5 |
| Cd1—S1—Cd1i | 125.12 (3) | ||
| N1ii—Cd1—S1—C1 | 30.55 (13) | N2—N1—C1—N4 | 0.2 (3) |
| N1i—Cd1—S1—C1 | −149.45 (13) | Cd1iii—N1—C1—N4 | 167.5 (2) |
| S1iii—Cd1—S1—C1 | −62.75 (11) | N2—N1—C1—S1 | 179.1 (2) |
| S1v—Cd1—S1—C1 | 117.25 (11) | Cd1iii—N1—C1—S1 | −13.6 (5) |
| N1ii—Cd1—S1—Cd1i | 163.99 (5) | N3—N4—C1—N1 | −0.4 (3) |
| N1i—Cd1—S1—Cd1i | −16.01 (5) | C2—N4—C1—N1 | −173.6 (3) |
| S1iii—Cd1—S1—Cd1i | 70.68 (4) | N3—N4—C1—S1 | −179.4 (2) |
| S1v—Cd1—S1—Cd1i | −109.32 (4) | C2—N4—C1—S1 | 7.4 (5) |
| C1—N1—N2—N3 | 0.0 (3) | Cd1—S1—C1—N1 | 64.1 (3) |
| Cd1iii—N1—N2—N3 | −171.82 (17) | Cd1i—S1—C1—N1 | −76.9 (3) |
| N1—N2—N3—N4 | −0.2 (3) | Cd1—S1—C1—N4 | −117.1 (2) |
| N2—N3—N4—C1 | 0.4 (3) | Cd1i—S1—C1—N4 | 101.9 (3) |
| N2—N3—N4—C2 | 174.3 (3) |
| Symmetry codes: (i) −x+1, y+1/2, −z+1/2; (ii) x, −y+1/2, z+1/2; (iii) −x+1, y−1/2, −z+1/2; (iv) −x+1, −y+1, −z+1; (v) x, −y+3/2, z+1/2. |

 journal menu
journal menu










 Alert level B
Alert level B
 Alert level C
Alert level C



