issue contents
June 2021 issue
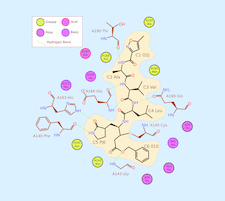
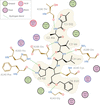
Cover illustration: The importance of correctly modelling covalent linkages is demonstrated using a case study which involves the covalent linkage of an inhibitor to the main protease (MPro) of SARS-CoV-2 [Nicholls et al. (2021), Acta Cryst. D77, 727–745]. This example demonstrates the importance of properly modelling covalent linkages using a comprehensive restraint dictionary, as opposed to just using a single interatomic distance restraint or failing to model the covalent linkage at all. This interaction analysis of N3 binding to SARS-CoV-2 MPro (PDB entry 6lu7) was created using Inkscape based on a template generated using LigPlot.
CCP4
Open  access
access
 access
accessThe mechanism for modelling covalent linkages in CCP4 is reviewed and the method of link-dictionary generation used by AceDRG is described. An overview of the various protocols available for the modelling and application of covalent linkages within the CCP4 suite is presented, providing instructive guidelines with a focus on practical application.
Open  access
access
 access
accessAnalysis of the Protein Data Bank revealed that over a third of entries contain covalent linkages without descriptions in the CCP4 Monomer Library (CCP4-ML). The CCP4-ML was updated with AceDRG dictionaries corresponding to commonly occurring classes of missing linkages.
Open  access
access
 access
accessVisualization renders structural molecular data accessible to a broad audience. An approach to share molecular-visualization experiences based on FAIR principles is described. The workflow is exemplified with recent COVID-19-related data.
research papers
Open  access
access
 access
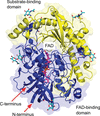
accessThe high-resolution crystal structure of a novel FAD-dependent oxidoreductase from the GMC oxidoreductase superfamily reveals a novel His–Ser active-site pair located in an extensive active-site pocket. Crystallographic fragment screening led to the identification of subsites inside the active-site pocket, indicating a preference for polyaromatic substrates.
PDB references: CtFDO, 6ze2; complex with (4-methoxycarbonylphenyl)methylazanium, 6ze3; complex with 4-oxo-N-[(1S)-1-(pyridin-3-yl)ethyl]-4-(thiophen-2-yl)butanamide, 6ze4; complex with 2-(1H-indol-3-yl)-N-[(1-methyl-1H-pyrrol-2-yl)methyl]ethanamine, 6ze5; complex with 4-nitrocatechol, 6ze6; complex with 4-nitrophenol, 6ze7; complex with ABTS, 7aa2
Structural and mutational studies of the active-site residues and also of those forming salt bridges between subdomains and active-site residues of the adenylation domain were carried out in M. tuberculosis NAD+-dependent DNA ligase A. The results suggest that specific salt bridges mediated by Glu22, Glu26 and Glu87 are important for the correct subdomain geometry and that their disruption critically impacts on the adenylation activity. At the same time, even conservative mutations of His236 and Lys123 in the active site lead to severe activity loss.
SASBDB references: AdD–NMN complex, SASDEU5; AdD–NAD+ complex, SASDEC6; AdD–NMN–AMP complex, SASDG65
Open  access
access
 access
accessTwo modulated structures of the Hyp-1–ANS complex with different unit-cell dimensions and contents were analyzed in (3+1)D superspace, revealing that they are very similar if not the same higher-dimensional structure with a slight shift in the q vector being responsible for the differences observed in 3D space.
Open  access
access
 access
accessThe amount of data generated during crystallographic fragment-screening projects requires sophisticated automated methods to analyse the data and to identify binders. FragMAXapp is the MAX IV Laboratory approach to managing fragment-screening campaigns: a web application that provides scientists with access to the MAX IV computing cluster and visualization tools.
Blood coagulation factor VIIa contains sodium, magnesium and calcium sites. In the present study, sodium increased the amidolytic activity of factor VIIa by ∼20-fold and stabilized the sodium loops and the tissue factor-binding region; however, rubidium occupied two calcium sites in factor VIIa but not the sodium site.
PDB reference: factor VIIa–soluble tissue factor complex, 4ibl
Open  access
access
 access
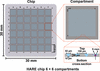
accessThe in-chip crystallization and structure determination of four different soluble proteins and an intracellular protein using serial synchrotron crystallography is reported.
Open  access
access
 access
accessPrincipal component analysis can only give a low-frequency description of the movements that a macromolecule is undergoing.
Open  access
access
 access
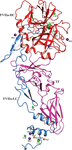
accessRsm22-family proteins are conserved putative SAM-dependent methyltransferases with important functions in mitochondrial translation. Here, the results of a comparative bioinformatics analysis of Rsm22-type proteins are presented, the expression, biophysical characterization and crystallization of Saccharomyces cerevisiae Rsm22 are reported, a low-resolution SAXS structure of the protein is revealed, and SAM-dependent RNA methyl transferase activity of the protein is demonstrated.
Open  access
access
 access
accessThe crystal structures of three viral Src kinase (v-Src) SH3-domain variants have been determined in order to obtain insights into the roles of these oncogenic mutations in the function of v-Src.


 journal menu
journal menu