issue contents
April 2005 issue
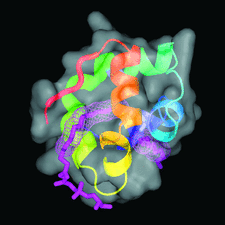
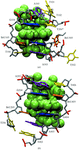
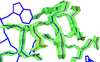
Cover illustration: Model of the non-specific type 2 lipid transfer protein from wheat, liganded with L-![[alpha]](/logos/entities/alpha_rmgif.gif) -palmitoyl-phosphatidyl glycerol (p. 397).
-palmitoyl-phosphatidyl glycerol (p. 397).
research papers
The 1.9 Å native structure of the pyrimidine-biosynthesis regulatory protein encoded by the M. tuberculosis pyrR gene (Rv1379) is reported. Because pyrimidine biosynthesis is an essential step in the progression of TB, pyrR is an attractive antitubercular drug target.
PDB reference: pyrR, 1w30, r1w30sf
The structure of CDP-D-glucose 4,6-dehydratase from S. typhi complexed with CDP-D-xylose has been determined to 1.8 Å resolution.
PDB reference: CDP-D-glucose 4,6-dehydratase, 1wvg, r1wvgsf
This paper reports the first atomic structure of a bacterial sorbitol dehydrogenase, with a model of the active site containing the cofactor NAD and the substate sorbitol.
Open  access
access
 access
accessA multi-subunit mitochondrial membrane-protein complex involved in the Krebs cycle and respiratory chain has been crystallized in a form suitable for near-atomic resolution structure determination.
The routine use of the single-wavelength anomalous diffraction technique (SAD) at wavelengths remote from absorption edges as a method for automated structure determination is discussed.
The crystal structure of wheat ns-LTP2 was determined ab initio to 1.12 Å resolution by direct methods. The α-helical bundle fold of this protein is maintained by four disulfide bridges and delineates two hydrophobic cavities filled by the alkyl chains of lipids.
PDB reference: wheat ns-LTP2, 1tuk, r1tuksf
In the crystal, actinomycin D intercalates between G–T wobble pairs of DNA involving the sequence d(TTAGTT); in solution, the dominant binding is between guanine and actinomycin.
A human–mouse chimeric antibody lacking any CH2 domain has been crystallized for diffraction analysis in the presence of detergent. Analysis by atomic force microscopy reveals the symmetry properties of the asymmetric unit and the presence of an unusual kind of defect within the crystals.
Powder diffraction data are shown to be sufficient to position a molecule of turkey egg-white lysozyme in the hexagonal unit cell by molecular replacement and to refine the structure at 3.35 Å resolution. The variation of the structure with sample preparation and upon freezing are also reported.
PDB reference: turkey egg-white lysozyme, 1xft, r1xftsf
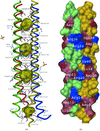
A 1.14 Å resolution study on the gramicidin D complex with RbCl confirms its right-handed antiparallel double-stranded dimeric structure. Rb cations distributed over seven binding sites in each gramicidin channel unit are `coordinated' by delocalized π-electrons of three to five carbonyl groups from the channel wall and by two water molecules.
PDB reference: gramicidin D–RbCl complex, 1w5u, r1w5usf
The classical bis-intercalation complexes of echinomycin with DNA have been examined at high resolution.
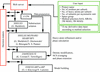
A description of the EMBL-Hamburg Automated Crystal Structure Determination Platform is given and examples of successful structure determination are presented.
Open  access
access
 access
accessDerivation of likelihood-enhanced translation functions that can be calculated by fast Fourier transform.
TLSView is an interactive molecular viewer for the generation, visualization and analysis of TLS (translation/libration/screw) models of protein motion.
The structure of a pyridoxal phosphate-dependent catalytic antibody in complex with a substrate analogue has been determined from a monoclinic pseudomerohedrally twinned crystal. Iodide molecules are present in the combining site and at intermolecular contacts.
PDB reference: 15A9–PPL-L-Ala complex, 1wc7
short communications
A novel approach for optimizing crystal contacts in a most favourable fashion by performing crystallization setups with the zwitterionic surfactant cocoamidoproylbetaine (CAPB) is introduced.
PDB reference: RHCC–CAPB, 1ybk, r1ybksf
The crystal structure of the S35C flavodoxin from D. vulgaris in the semiquinone state at 1.8 Å resolution. shows as main difference with the oxidized form a conformational change of the carbonyl oxygen of Gly61, leading to the formation of a new hydrogen bond between the protein and the prosthetic group.
PDB reference: S35C flavodoxin mutant, 1xt6, r1xt6sf
The clottable protein from the California spiny lobster has been purified and crystallized in two forms. One of the crystal forms exhibits unusual pseudo-ninefold rotational symmetry in space group P21.
Equilibrating crystallization drops against different reservoirs can alter the results of a vapor-diffusion crystallization experiment.


 journal menu
journal menu