The title compound, crystallized with two conformationally similar molecules (
A and
B) in the asymmetric unit. In the crystal, individual molecules are linked by pairs of O—H

O hydrogen bonds, forming
A–
A and
B–
B inversion dimers with
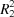
(12) ring motifs.
Supporting information
CCDC reference: 1009715
Key indicators
- Single-crystal X-ray study
- T = 173 K
- Mean
![[sigma]](https://journals.iucr.org/logos/entities/sigma_rmgif.gif) (C-C) = 0.003 Å
(C-C) = 0.003 Å
- R factor = 0.030
- wR factor = 0.078
- Data-to-parameter ratio = 13.5
checkCIF/PLATON results
No syntax errors found
 Alert level C
PLAT480_ALERT_4_C Long H...A H-Bond Reported H6 .. CL4 .. 2.91 Ang.
PLAT906_ALERT_3_C Large K value in the Analysis of Variance ...... 2.244 Check
PLAT911_ALERT_3_C Missing # FCF Refl Between THmin & STh/L= 0.600 8 Report
PLAT913_ALERT_3_C Missing # of Very Strong Reflections in FCF .... 1 Note
Alert level C
PLAT480_ALERT_4_C Long H...A H-Bond Reported H6 .. CL4 .. 2.91 Ang.
PLAT906_ALERT_3_C Large K value in the Analysis of Variance ...... 2.244 Check
PLAT911_ALERT_3_C Missing # FCF Refl Between THmin & STh/L= 0.600 8 Report
PLAT913_ALERT_3_C Missing # of Very Strong Reflections in FCF .... 1 Note
 Alert level G
PLAT793_ALERT_4_G The Model has Chirality at C2 (Centro SPGR) R Verify
PLAT793_ALERT_4_G The Model has Chirality at C5 (Centro SPGR) R Verify
PLAT912_ALERT_4_G Missing # of FCF Reflections Above STh/L= 0.600 10 Note
Alert level G
PLAT793_ALERT_4_G The Model has Chirality at C2 (Centro SPGR) R Verify
PLAT793_ALERT_4_G The Model has Chirality at C5 (Centro SPGR) R Verify
PLAT912_ALERT_4_G Missing # of FCF Reflections Above STh/L= 0.600 10 Note
0 ALERT level A = Most likely a serious problem - resolve or explain
0 ALERT level B = A potentially serious problem, consider carefully
4 ALERT level C = Check. Ensure it is not caused by an omission or oversight
3 ALERT level G = General information/check it is not something unexpected
0 ALERT type 1 CIF construction/syntax error, inconsistent or missing data
0 ALERT type 2 Indicator that the structure model may be wrong or deficient
3 ALERT type 3 Indicator that the structure quality may be low
4 ALERT type 4 Improvement, methodology, query or suggestion
0 ALERT type 5 Informative message, check
Data collection: X-AREA (Stoe & Cie, 2009); cell refinement: X-AREA (Stoe & Cie, 2009); data reduction: X-RED32 (Stoe & Cie, 2009); program(s) used to solve structure: SHELXS2014 (Sheldrick, 2008); program(s) used to refine structure: SHELXL2014 (Sheldrick, 2015); molecular graphics: PLATON (Spek, 2009) and Mercury (Macrae et al., 2008); software used to prepare material for publication: SHELXL2014 (Sheldrick, 2015) and PLATON (Spek, 2009).
N-(2,2,2-Trichloro-1-hydroxyethyl)formamide
top
Crystal data top
| C3H4Cl3NO2 | F(000) = 768 |
| Mr = 192.42 | Dx = 1.830 Mg m−3 |
| Monoclinic, P21/c | Mo Kα radiation, λ = 0.71073 Å |
| a = 13.7964 (8) Å | Cell parameters from 22309 reflections |
| b = 9.0798 (7) Å | θ = 1.6–26.2° |
| c = 12.2453 (7) Å | µ = 1.24 mm−1 |
| β = 114.413 (4)° | T = 173 K |
| V = 1396.80 (16) Å3 | Block, colourless |
| Z = 8 | 0.45 × 0.43 × 0.40 mm |
Data collection top
Stoe IPDS 2
diffractometer | 2645 independent reflections |
| Radiation source: fine-focus sealed tube | 2468 reflections with I > 2σ(I) |
| Plane graphite monochromator | Rint = 0.056 |
| φ + ω scans | θmax = 25.7°, θmin = 1.6° |
Absorption correction: multi-scan
(MULABS in PLATON; Spek, 2009) | h = −15→16 |
| Tmin = 0.579, Tmax = 1.000 | k = −11→11 |
| 16347 measured reflections | l = −14→14 |
Refinement top
| Refinement on F2 | Hydrogen site location: difference Fourier map |
| Least-squares matrix: full | All H-atom parameters refined |
| R[F2 > 2σ(F2)] = 0.030 | w = 1/[σ2(Fo2) + (0.0333P)2 + 1.2964P]
where P = (Fo2 + 2Fc2)/3 |
| wR(F2) = 0.078 | (Δ/σ)max < 0.001 |
| S = 1.08 | Δρmax = 0.82 e Å−3 |
| 2645 reflections | Δρmin = −0.55 e Å−3 |
| 196 parameters | Extinction correction: SHELXL2014 (Sheldrick, 2015), Fc*=kFc[1+0.001xFc2λ3/sin(2θ)]-1/4 |
| 0 restraints | Extinction coefficient: 0.0089 (9) |
Special details top
Geometry. All e.s.d.'s (except the e.s.d. in the dihedral angle between two l.s. planes)
are estimated using the full covariance matrix. The cell e.s.d.'s are taken
into account individually in the estimation of e.s.d.'s in distances, angles
and torsion angles; correlations between e.s.d.'s in cell parameters are only
used when they are defined by crystal symmetry. An approximate (isotropic)
treatment of cell e.s.d.'s is used for estimating e.s.d.'s involving l.s.
planes. |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2) top| | x | y | z | Uiso*/Ueq | |
| Cl1 | 0.13449 (5) | 0.04880 (6) | 0.08600 (5) | 0.03428 (17) | |
| Cl2 | 0.22308 (4) | 0.28605 (6) | −0.00018 (5) | 0.03123 (16) | |
| Cl3 | 0.28102 (5) | 0.25302 (8) | 0.25345 (5) | 0.04298 (19) | |
| O1 | −0.00275 (12) | 0.31695 (18) | −0.01499 (14) | 0.0284 (4) | |
| H1O | 0.005 (3) | 0.366 (4) | −0.069 (3) | 0.058 (10)* | |
| O2 | 0.00144 (13) | 0.53996 (16) | 0.21058 (14) | 0.0295 (4) | |
| N1 | 0.04737 (14) | 0.30393 (19) | 0.18967 (16) | 0.0213 (4) | |
| H1N | 0.0392 (18) | 0.215 (3) | 0.207 (2) | 0.020 (6)* | |
| C1 | 0.17653 (17) | 0.2341 (2) | 0.10963 (19) | 0.0234 (4) | |
| C2 | 0.08245 (16) | 0.3364 (2) | 0.09670 (18) | 0.0217 (4) | |
| H2 | 0.1107 (18) | 0.434 (3) | 0.1068 (19) | 0.020 (6)* | |
| C3 | 0.00885 (17) | 0.4079 (2) | 0.23751 (18) | 0.0237 (4) | |
| H3 | −0.0156 (18) | 0.374 (3) | 0.299 (2) | 0.026 (6)* | |
| Cl4 | 0.64204 (5) | 0.45558 (5) | 0.52480 (5) | 0.03177 (16) | |
| Cl5 | 0.78172 (4) | 0.22393 (6) | 0.51229 (5) | 0.02994 (15) | |
| Cl6 | 0.71719 (4) | 0.23240 (7) | 0.70803 (5) | 0.03280 (16) | |
| O3 | 0.49315 (12) | 0.20023 (18) | 0.51142 (15) | 0.0274 (3) | |
| H3O | 0.497 (2) | 0.153 (3) | 0.564 (3) | 0.036 (8)* | |
| O4 | 0.50422 (13) | −0.04575 (15) | 0.29695 (13) | 0.0281 (3) | |
| N2 | 0.54593 (14) | 0.19299 (19) | 0.35510 (15) | 0.0208 (4) | |
| H2N | 0.5415 (19) | 0.274 (3) | 0.332 (2) | 0.023 (6)* | |
| C4 | 0.67565 (16) | 0.2671 (2) | 0.55274 (18) | 0.0216 (4) | |
| C5 | 0.57819 (16) | 0.1686 (2) | 0.48142 (18) | 0.0206 (4) | |
| H5 | 0.6025 (16) | 0.066 (2) | 0.5015 (18) | 0.012 (5)* | |
| C6 | 0.51071 (16) | 0.0854 (2) | 0.27421 (18) | 0.0230 (4) | |
| H6 | 0.4911 (19) | 0.119 (3) | 0.190 (2) | 0.028 (6)* | |
Atomic displacement parameters (Å2) top| | U11 | U22 | U33 | U12 | U13 | U23 |
| Cl1 | 0.0493 (4) | 0.0157 (3) | 0.0449 (3) | 0.0062 (2) | 0.0265 (3) | 0.0024 (2) |
| Cl2 | 0.0363 (3) | 0.0317 (3) | 0.0356 (3) | 0.0029 (2) | 0.0248 (2) | 0.0016 (2) |
| Cl3 | 0.0250 (3) | 0.0729 (5) | 0.0282 (3) | 0.0012 (3) | 0.0081 (2) | −0.0095 (3) |
| O1 | 0.0292 (8) | 0.0315 (9) | 0.0259 (8) | 0.0064 (6) | 0.0127 (7) | 0.0100 (7) |
| O2 | 0.0478 (10) | 0.0170 (7) | 0.0307 (8) | 0.0078 (6) | 0.0232 (7) | 0.0031 (6) |
| N1 | 0.0268 (9) | 0.0138 (8) | 0.0279 (9) | 0.0012 (7) | 0.0159 (7) | 0.0025 (7) |
| C1 | 0.0259 (10) | 0.0227 (10) | 0.0236 (10) | 0.0014 (8) | 0.0121 (9) | −0.0014 (8) |
| C2 | 0.0272 (10) | 0.0144 (10) | 0.0275 (10) | 0.0016 (8) | 0.0154 (8) | 0.0010 (8) |
| C3 | 0.0288 (11) | 0.0221 (10) | 0.0229 (10) | 0.0034 (8) | 0.0133 (9) | 0.0013 (8) |
| Cl4 | 0.0415 (3) | 0.0152 (2) | 0.0346 (3) | −0.0028 (2) | 0.0117 (2) | −0.0021 (2) |
| Cl5 | 0.0247 (3) | 0.0366 (3) | 0.0304 (3) | 0.0009 (2) | 0.0133 (2) | 0.0020 (2) |
| Cl6 | 0.0336 (3) | 0.0425 (3) | 0.0191 (3) | −0.0029 (2) | 0.0077 (2) | 0.0028 (2) |
| O3 | 0.0287 (8) | 0.0303 (8) | 0.0265 (8) | −0.0020 (6) | 0.0150 (7) | 0.0051 (7) |
| O4 | 0.0426 (9) | 0.0170 (7) | 0.0273 (8) | −0.0055 (6) | 0.0171 (7) | −0.0019 (6) |
| N2 | 0.0271 (9) | 0.0132 (8) | 0.0202 (9) | −0.0017 (7) | 0.0080 (7) | 0.0025 (7) |
| C4 | 0.0253 (10) | 0.0199 (10) | 0.0199 (10) | −0.0008 (8) | 0.0096 (8) | 0.0008 (7) |
| C5 | 0.0253 (10) | 0.0147 (10) | 0.0216 (10) | −0.0018 (8) | 0.0095 (8) | 0.0016 (7) |
| C6 | 0.0270 (10) | 0.0211 (10) | 0.0217 (10) | −0.0021 (8) | 0.0111 (8) | −0.0007 (8) |
Geometric parameters (Å, º) top
| Cl1—C1 | 1.764 (2) | Cl4—C4 | 1.769 (2) |
| Cl2—C1 | 1.777 (2) | Cl5—C4 | 1.772 (2) |
| Cl3—C1 | 1.764 (2) | Cl6—C4 | 1.773 (2) |
| O1—C2 | 1.398 (3) | O3—C5 | 1.396 (3) |
| O1—H1O | 0.84 (3) | O3—H3O | 0.76 (3) |
| O2—C3 | 1.236 (3) | O4—C6 | 1.235 (3) |
| N1—C3 | 1.332 (3) | N2—C6 | 1.332 (3) |
| N1—C2 | 1.440 (3) | N2—C5 | 1.439 (3) |
| N1—H1N | 0.85 (2) | N2—H2N | 0.78 (3) |
| C1—C2 | 1.550 (3) | C4—C5 | 1.549 (3) |
| C2—H2 | 0.96 (2) | C5—H5 | 0.99 (2) |
| C3—H3 | 0.99 (2) | C6—H6 | 1.00 (2) |
| | | |
| C2—O1—H1O | 112 (2) | C5—O3—H3O | 110 (2) |
| C3—N1—C2 | 121.88 (17) | C6—N2—C5 | 122.84 (17) |
| C3—N1—H1N | 116.3 (16) | C6—N2—H2N | 118.0 (18) |
| C2—N1—H1N | 120.8 (16) | C5—N2—H2N | 118.7 (18) |
| C2—C1—Cl3 | 110.37 (14) | C5—C4—Cl4 | 110.64 (14) |
| C2—C1—Cl1 | 110.49 (14) | C5—C4—Cl5 | 109.84 (14) |
| Cl3—C1—Cl1 | 109.52 (11) | Cl4—C4—Cl5 | 109.91 (11) |
| C2—C1—Cl2 | 108.24 (14) | C5—C4—Cl6 | 108.82 (14) |
| Cl3—C1—Cl2 | 109.15 (11) | Cl4—C4—Cl6 | 108.79 (11) |
| Cl1—C1—Cl2 | 109.04 (11) | Cl5—C4—Cl6 | 108.80 (11) |
| O1—C2—N1 | 109.04 (17) | O3—C5—N2 | 109.32 (16) |
| O1—C2—C1 | 110.74 (16) | O3—C5—C4 | 111.15 (16) |
| N1—C2—C1 | 109.73 (16) | N2—C5—C4 | 109.14 (16) |
| O1—C2—H2 | 112.1 (13) | O3—C5—H5 | 111.4 (12) |
| N1—C2—H2 | 109.8 (14) | N2—C5—H5 | 109.6 (12) |
| C1—C2—H2 | 105.3 (14) | C4—C5—H5 | 106.2 (12) |
| O2—C3—N1 | 125.03 (19) | O4—C6—N2 | 125.30 (19) |
| O2—C3—H3 | 119.1 (14) | O4—C6—H6 | 120.9 (14) |
| N1—C3—H3 | 115.9 (14) | N2—C6—H6 | 113.7 (14) |
| | | |
| C3—N1—C2—O1 | −91.5 (2) | C6—N2—C5—O3 | −95.7 (2) |
| C3—N1—C2—C1 | 147.08 (19) | C6—N2—C5—C4 | 142.52 (19) |
| Cl3—C1—C2—O1 | −177.16 (14) | Cl4—C4—C5—O3 | −58.44 (19) |
| Cl1—C1—C2—O1 | −55.9 (2) | Cl5—C4—C5—O3 | −179.96 (13) |
| Cl2—C1—C2—O1 | 63.45 (19) | Cl6—C4—C5—O3 | 61.03 (19) |
| Cl3—C1—C2—N1 | −56.7 (2) | Cl4—C4—C5—N2 | 62.20 (19) |
| Cl1—C1—C2—N1 | 64.54 (19) | Cl5—C4—C5—N2 | −59.31 (19) |
| Cl2—C1—C2—N1 | −176.13 (14) | Cl6—C4—C5—N2 | −178.33 (14) |
| C2—N1—C3—O2 | −1.8 (3) | C5—N2—C6—O4 | −2.1 (3) |
Hydrogen-bond geometry (Å, º) top
| D—H···A | D—H | H···A | D···A | D—H···A |
| O1—H1O···O2i | 0.84 (3) | 1.90 (4) | 2.731 (2) | 169 (3) |
| N1—H1N···O2ii | 0.85 (2) | 2.08 (3) | 2.893 (2) | 159 (2) |
| O3—H3O···O4iii | 0.76 (3) | 1.97 (3) | 2.721 (2) | 174 (3) |
| N2—H2N···O4iv | 0.78 (3) | 2.17 (3) | 2.917 (2) | 158 (2) |
| C6—H6···Cl4v | 1.00 (2) | 2.91 (2) | 3.586 (2) | 125 (2) |
| Symmetry codes: (i) −x, −y+1, −z; (ii) −x, y−1/2, −z+1/2; (iii) −x+1, −y, −z+1; (iv) −x+1, y+1/2, −z+1/2; (v) −x+1, y−1/2, −z+1/2. |
 access
access O hydrogen bonds, forming A–A and B–B inversion dimers with
O hydrogen bonds, forming A–A and B–B inversion dimers with ![[sigma]](https://journals.iucr.org/logos/entities/sigma_rmgif.gif) (C-C) = 0.003 Å
(C-C) = 0.003 ÅAlert level C PLAT480_ALERT_4_C Long H...A H-Bond Reported H6 .. CL4 .. 2.91 Ang. PLAT906_ALERT_3_C Large K value in the Analysis of Variance ...... 2.244 Check PLAT911_ALERT_3_C Missing # FCF Refl Between THmin & STh/L= 0.600 8 Report PLAT913_ALERT_3_C Missing # of Very Strong Reflections in FCF .... 1 Note
Alert level G PLAT793_ALERT_4_G The Model has Chirality at C2 (Centro SPGR) R Verify PLAT793_ALERT_4_G The Model has Chirality at C5 (Centro SPGR) R Verify PLAT912_ALERT_4_G Missing # of FCF Reflections Above STh/L= 0.600 10 Note

 journal menu
journal menu











