The title compound, C
24H
23ClN
4, crystallizes in space group
P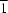
, with two crystallographically independent molecules in the asymmetric unit. In both molecules, the pyrrolidine ring adopts a half-chair conformation. The molecular packing in the crystal is stabilized by intermolecular C—H

N hydrogen bonds, through the formation of hydrogen-bonded molecular chains.
Supporting information
CCDC reference: 202362
Key indicators
- Single-crystal X-ray study
- T = 293 K
- Mean
![[sigma]](//journals.iucr.org/logos/entities/sigma_rmgif.gif) (C-C) = 0.003 Å
(C-C) = 0.003 Å
- R factor = 0.054
- wR factor = 0.160
- Data-to-parameter ratio = 18.2
checkCIF results
No syntax errors found
ADDSYM reports no extra symmetry
 Alert Level C:
Alert Level C:
REFLT_03
From the CIF: _diffrn_reflns_theta_max 28.31
From the CIF: _reflns_number_total 9579
TEST2: Reflns within _diffrn_reflns_theta_max
Count of symmetry unique reflns 10436
Completeness (_total/calc) 91.79%
Alert C: < 95% complete
PLAT_360 Alert C Short C(sp3)-C(sp3) Bond C(8B) - C(9B) = 1.42 Ang.
PLAT_371 Alert C Long C(sp2)-C(sp1) Bond C(3A) - C(11A) = 1.43 Ang.
PLAT_371 Alert C Long C(sp2)-C(sp1) Bond C(3B) - C(11B) = 1.44 Ang.
0 Alert Level A = Potentially serious problem
0 Alert Level B = Potential problem
4 Alert Level C = Please check
To a refluxing solution of 4-dimethylamino-4'-(chlorobenzoyl)acetophenone (1.8 mmol) in 10 ml of ethanol, malononitrile (1.8 mmol) and pyrrolidine (1.8 mmol) were added and the resulting solution refluxed for 9 h. The solvent was distilled off under reduced pressure and the resulting residue was purified by column chromatography on silica-gel (100–200 mesh). Single crystals were obtained by slow evaporation method using petroleum ether and ethyl acetate (1:3) solvent system.
All the H atoms were fixed geometrically and allowed to ride on the corresponding non-H atoms, with C—H distances of 0.93–0.97 Å, and Uiso(H) = 1.5Ueq(C) for methyl H atoms and 1.2Ueq(C) for all other H atoms. Rotating-group refinement was used for the methyl H atoms. Large displacement parameters for atom C9B indicate the possibility of conformational disorder for the pyrrolidine ring. This resulted in a shortening of the C8B—C9B [1.416 (4) Å] bond. Attempts to refine the structure using a disorder model for the pyrrolidine ring resulted in unrealistic Csp3—Csp3 distances and hence the original model was retained.
Data collection: SMART (Siemens, 1996); cell refinement: SAINT (Siemens, 1996); data reduction: SAINT; program(s) used to solve structure: SHELXS97 (Sheldrick, 1997); program(s) used to refine structure: SHELXL97 (Sheldrick, 1997); molecular graphics: ZORTEP (Zsolnai, 1997); software used to prepare material for publication: SHELXL97 and PARST (Nardelli, 1983, 1995).
6-(4-Chlorophenyl)-4-[4-(dimethylamino)phenyl]-2-(pyrrolidin-1- yl)nicotinonitrile
top
Crystal data top
| C24H23ClN4 | Z = 4 |
| Mr = 402.91 | F(000) = 848 |
| Triclinic, P1 | Dx = 1.276 Mg m−3 |
| Hall symbol: -P 1 | Mo Kα radiation, λ = 0.71073 Å |
| a = 11.1490 (1) Å | Cell parameters from 7711 reflections |
| b = 13.7330 (1) Å | θ = 1.4–28.3° |
| c = 14.6666 (1) Å | µ = 0.20 mm−1 |
| α = 100.487 (1)° | T = 293 K |
| β = 93.240 (1)° | Block, yellow |
| γ = 107.014 (1)° | 0.48 × 0.44 × 0.26 mm |
| V = 2096.93 (2) Å3 | |
Data collection top
Siemens SMART CCD area-detector
diffractometer | 6307 reflections with I > 2σ(I) |
| Radiation source: fine-focus sealed tube | Rint = 0.048 |
| Graphite monochromator | θmax = 28.3°, θmin = 1.4° |
| ω scans | h = −13→14 |
| 14581 measured reflections | k = −17→14 |
| 9579 independent reflections | l = −12→19 |
Refinement top
| Refinement on F2 | Primary atom site location: structure-invariant direct methods |
| Least-squares matrix: full | Secondary atom site location: difference Fourier map |
| R[F2 > 2σ(F2)] = 0.054 | Hydrogen site location: inferred from neighbouring sites |
| wR(F2) = 0.160 | H-atom parameters constrained |
| S = 0.97 | w = 1/[σ2(Fo2) + (0.0793P)2]
where P = (Fo2 + 2Fc2)/3 |
| 9579 reflections | (Δ/σ)max < 0.001 |
| 527 parameters | Δρmax = 0.35 e Å−3 |
| 0 restraints | Δρmin = −0.32 e Å−3 |
Crystal data top
| C24H23ClN4 | γ = 107.014 (1)° |
| Mr = 402.91 | V = 2096.93 (2) Å3 |
| Triclinic, P1 | Z = 4 |
| a = 11.1490 (1) Å | Mo Kα radiation |
| b = 13.7330 (1) Å | µ = 0.20 mm−1 |
| c = 14.6666 (1) Å | T = 293 K |
| α = 100.487 (1)° | 0.48 × 0.44 × 0.26 mm |
| β = 93.240 (1)° | |
Data collection top
Siemens SMART CCD area-detector
diffractometer | 6307 reflections with I > 2σ(I) |
| 14581 measured reflections | Rint = 0.048 |
| 9579 independent reflections | |
Refinement top
| R[F2 > 2σ(F2)] = 0.054 | 0 restraints |
| wR(F2) = 0.160 | H-atom parameters constrained |
| S = 0.97 | Δρmax = 0.35 e Å−3 |
| 9579 reflections | Δρmin = −0.32 e Å−3 |
| 527 parameters | |
Special details top
Geometry. All e.s.d.'s (except the e.s.d. in the dihedral angle between two l.s. planes) are estimated using the full covariance matrix. The cell e.s.d.'s are taken into account individually in the estimation of e.s.d.'s in distances, angles and torsion angles; correlations between e.s.d.'s in cell parameters are only used when they are defined by crystal symmetry. An approximate (isotropic) treatment of cell e.s.d.'s is used for estimating e.s.d.'s involving l.s. planes. |
Refinement. Refinement of F2 against ALL reflections. The weighted R-factor wR and goodness of fit S are based on F2, conventional R-factors R are based on F, with F set to zero for negative F2. The threshold expression of F2 > σ(F2) is used only for calculating R-factors(gt) etc. and is not relevant to the choice of reflections for refinement. R-factors based on F2 are statistically about twice as large as those based on F, and R- factors based on ALL data will be even larger. |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2) top| | x | y | z | Uiso*/Ueq | |
| Cl1A | −0.02596 (6) | −0.38678 (5) | 0.59338 (5) | 0.0866 (2) | |
| N1A | 0.21081 (13) | 0.08918 (11) | 0.51050 (9) | 0.0431 (3) | |
| N2A | 0.28907 (15) | 0.26496 (11) | 0.55895 (10) | 0.0481 (4) | |
| N3A | 0.39956 (18) | 0.35530 (13) | 0.34391 (12) | 0.0628 (4) | |
| N4A | 0.15588 (17) | 0.09242 (14) | −0.06908 (11) | 0.0640 (5) | |
| C2A | 0.25494 (16) | 0.18186 (13) | 0.48645 (11) | 0.0407 (4) | |
| C3A | 0.26408 (16) | 0.18899 (13) | 0.39066 (11) | 0.0407 (4) | |
| C4A | 0.21067 (16) | 0.09795 (13) | 0.32108 (11) | 0.0404 (4) | |
| C5A | 0.16151 (17) | 0.00392 (13) | 0.34922 (11) | 0.0437 (4) | |
| H5A | 0.1256 | −0.0573 | 0.3049 | 0.052* | |
| C6A | 0.16631 (15) | 0.00203 (13) | 0.44416 (11) | 0.0397 (4) | |
| C7A | 0.2726 (2) | 0.24839 (15) | 0.65427 (12) | 0.0577 (5) | |
| H7A | 0.3256 | 0.2089 | 0.6727 | 0.069* | |
| H7B | 0.1853 | 0.2116 | 0.6591 | 0.069* | |
| C8A | 0.3119 (3) | 0.35634 (18) | 0.71363 (16) | 0.0777 (7) | |
| H8A | 0.3994 | 0.3765 | 0.7404 | 0.093* | |
| H8B | 0.2595 | 0.3601 | 0.7639 | 0.093* | |
| C9A | 0.2948 (2) | 0.42604 (16) | 0.64905 (15) | 0.0685 (6) | |
| H9A | 0.3542 | 0.4954 | 0.6696 | 0.082* | |
| H9B | 0.2096 | 0.4309 | 0.6464 | 0.082* | |
| C10A | 0.3198 (2) | 0.37435 (14) | 0.55470 (13) | 0.0545 (5) | |
| H10A | 0.2665 | 0.3840 | 0.5047 | 0.065* | |
| H10B | 0.4075 | 0.4024 | 0.5450 | 0.065* | |
| C11A | 0.33675 (18) | 0.28231 (15) | 0.36491 (12) | 0.0470 (4) | |
| C12A | 0.20264 (16) | 0.10013 (13) | 0.22023 (11) | 0.0425 (4) | |
| C13A | 0.16198 (17) | 0.17451 (14) | 0.18509 (12) | 0.0459 (4) | |
| H13A | 0.1461 | 0.2278 | 0.2266 | 0.055* | |
| C14A | 0.14480 (17) | 0.17124 (15) | 0.09087 (13) | 0.0495 (4) | |
| H14A | 0.1159 | 0.2214 | 0.0703 | 0.059* | |
| C15A | 0.16985 (16) | 0.09399 (15) | 0.02500 (12) | 0.0472 (4) | |
| C16A | 0.21283 (19) | 0.02003 (15) | 0.06020 (12) | 0.0517 (4) | |
| H16A | 0.2322 | −0.0317 | 0.0190 | 0.062* | |
| C17A | 0.22686 (18) | 0.02288 (14) | 0.15493 (12) | 0.0500 (4) | |
| H17A | 0.2533 | −0.0284 | 0.1758 | 0.060* | |
| C18A | 0.11896 (15) | −0.09523 (13) | 0.47940 (11) | 0.0400 (4) | |
| C19A | 0.09205 (18) | −0.19282 (14) | 0.42202 (13) | 0.0488 (4) | |
| H19A | 0.1034 | −0.1980 | 0.3592 | 0.059* | |
| C20A | 0.04842 (19) | −0.28293 (15) | 0.45645 (14) | 0.0559 (5) | |
| H20A | 0.0314 | −0.3480 | 0.4173 | 0.067* | |
| C21A | 0.03079 (18) | −0.27462 (15) | 0.54923 (14) | 0.0538 (5) | |
| C22A | 0.05682 (19) | −0.17883 (16) | 0.60837 (14) | 0.0566 (5) | |
| H22A | 0.0450 | −0.1742 | 0.6711 | 0.068* | |
| C23A | 0.10068 (18) | −0.08987 (15) | 0.57333 (12) | 0.0493 (4) | |
| H23A | 0.1184 | −0.0251 | 0.6130 | 0.059* | |
| C24A | 0.1804 (3) | 0.0111 (2) | −0.13572 (15) | 0.0832 (7) | |
| H24A | 0.2623 | 0.0063 | −0.1181 | 0.125* | |
| H24B | 0.1777 | 0.0272 | −0.1967 | 0.125* | |
| H24C | 0.1174 | −0.0540 | −0.1366 | 0.125* | |
| C25A | 0.0992 (2) | 0.1623 (2) | −0.10475 (16) | 0.0761 (7) | |
| H25A | 0.1477 | 0.2330 | −0.0777 | 0.114* | |
| H25B | 0.0144 | 0.1495 | −0.0888 | 0.114* | |
| H25C | 0.0978 | 0.1507 | −0.1714 | 0.114* | |
| Cl1B | 0.56120 (7) | 0.11966 (5) | −0.06469 (5) | 0.0903 (2) | |
| N1B | 0.27692 (14) | 0.48400 (11) | 0.01936 (10) | 0.0450 (3) | |
| N2B | 0.16274 (16) | 0.58056 (13) | −0.03099 (10) | 0.0546 (4) | |
| N3B | 0.1058 (2) | 0.73715 (19) | 0.19077 (13) | 0.0944 (8) | |
| N4B | 0.40411 (19) | 0.80256 (14) | 0.58715 (12) | 0.0674 (5) | |
| C2B | 0.22383 (17) | 0.55999 (14) | 0.04215 (12) | 0.0448 (4) | |
| C3B | 0.23303 (17) | 0.61224 (14) | 0.13737 (12) | 0.0464 (4) | |
| C4B | 0.30233 (16) | 0.58518 (13) | 0.20667 (11) | 0.0427 (4) | |
| C5B | 0.35384 (16) | 0.50485 (13) | 0.18000 (12) | 0.0442 (4) | |
| H5B | 0.3966 | 0.4830 | 0.2246 | 0.053* | |
| C6B | 0.34082 (15) | 0.45748 (13) | 0.08610 (11) | 0.0407 (4) | |
| C7B | 0.1653 (2) | 0.52465 (18) | −0.12654 (13) | 0.0652 (6) | |
| H7C | 0.1231 | 0.4506 | −0.1342 | 0.078* | |
| H7D | 0.2513 | 0.5354 | −0.1413 | 0.078* | |
| C8B | 0.0948 (3) | 0.5730 (2) | −0.18689 (17) | 0.0961 (9) | |
| H8C | 0.1318 | 0.5785 | −0.2448 | 0.115* | |
| H8D | 0.0068 | 0.5311 | −0.2017 | 0.115* | |
| C9B | 0.1053 (4) | 0.6731 (3) | −0.13448 (19) | 0.1218 (14) | |
| H9C | 0.0323 | 0.6935 | −0.1517 | 0.146* | |
| H9D | 0.1805 | 0.7240 | −0.1465 | 0.146* | |
| C10B | 0.1131 (2) | 0.66733 (18) | −0.03218 (15) | 0.0681 (6) | |
| H10C | 0.1697 | 0.7313 | 0.0061 | 0.082* | |
| H10D | 0.0305 | 0.6537 | −0.0103 | 0.082* | |
| C11B | 0.1642 (2) | 0.68392 (17) | 0.16594 (13) | 0.0622 (5) | |
| C12B | 0.32554 (17) | 0.64203 (13) | 0.30520 (12) | 0.0435 (4) | |
| C13B | 0.37128 (18) | 0.74982 (14) | 0.32965 (13) | 0.0506 (4) | |
| H13B | 0.3836 | 0.7877 | 0.2826 | 0.061* | |
| C14B | 0.39903 (19) | 0.80240 (14) | 0.42143 (13) | 0.0523 (4) | |
| H14B | 0.4316 | 0.8746 | 0.4348 | 0.063* | |
| C15B | 0.37941 (17) | 0.74988 (14) | 0.49475 (12) | 0.0471 (4) | |
| C16B | 0.33469 (19) | 0.64076 (14) | 0.47045 (12) | 0.0521 (4) | |
| H16B | 0.3219 | 0.6028 | 0.5174 | 0.063* | |
| C17B | 0.30930 (18) | 0.58880 (14) | 0.37856 (12) | 0.0480 (4) | |
| H17B | 0.2806 | 0.5164 | 0.3649 | 0.058* | |
| C18B | 0.40065 (15) | 0.37612 (13) | 0.05188 (11) | 0.0401 (4) | |
| C19B | 0.44773 (18) | 0.32400 (15) | 0.11074 (13) | 0.0526 (4) | |
| H19B | 0.4461 | 0.3422 | 0.1748 | 0.063* | |
| C20B | 0.4970 (2) | 0.24541 (17) | 0.07529 (15) | 0.0608 (5) | |
| H20B | 0.5267 | 0.2099 | 0.1151 | 0.073* | |
| C21B | 0.50179 (18) | 0.22026 (15) | −0.01918 (15) | 0.0556 (5) | |
| C22B | 0.45904 (19) | 0.27182 (15) | −0.07922 (14) | 0.0555 (5) | |
| H22B | 0.4636 | 0.2547 | −0.1430 | 0.067* | |
| C23B | 0.40904 (18) | 0.34975 (14) | −0.04320 (12) | 0.0485 (4) | |
| H23B | 0.3804 | 0.3853 | −0.0835 | 0.058* | |
| C24B | 0.3554 (2) | 0.75155 (19) | 0.66077 (15) | 0.0736 (6) | |
| H24D | 0.2678 | 0.7128 | 0.6430 | 0.110* | |
| H24E | 0.3641 | 0.8028 | 0.7169 | 0.110* | |
| H24F | 0.4017 | 0.7050 | 0.6715 | 0.110* | |
| C25B | 0.4560 (3) | 0.91400 (17) | 0.60971 (17) | 0.0794 (7) | |
| H25D | 0.5295 | 0.9351 | 0.5783 | 0.119* | |
| H25E | 0.4789 | 0.9371 | 0.6759 | 0.119* | |
| H25F | 0.3941 | 0.9443 | 0.5900 | 0.119* | |
Atomic displacement parameters (Å2) top| | U11 | U22 | U33 | U12 | U13 | U23 |
| Cl1A | 0.0898 (5) | 0.0735 (4) | 0.0984 (5) | 0.0092 (3) | 0.0033 (3) | 0.0514 (3) |
| N1A | 0.0528 (8) | 0.0457 (8) | 0.0340 (7) | 0.0189 (6) | 0.0069 (6) | 0.0098 (6) |
| N2A | 0.0651 (9) | 0.0467 (8) | 0.0331 (7) | 0.0195 (7) | 0.0039 (6) | 0.0067 (6) |
| N3A | 0.0851 (12) | 0.0563 (10) | 0.0516 (10) | 0.0212 (9) | 0.0165 (9) | 0.0213 (8) |
| N4A | 0.0773 (12) | 0.0837 (12) | 0.0337 (8) | 0.0254 (10) | 0.0039 (8) | 0.0183 (8) |
| C2A | 0.0459 (9) | 0.0463 (9) | 0.0344 (8) | 0.0197 (7) | 0.0065 (7) | 0.0102 (7) |
| C3A | 0.0470 (9) | 0.0460 (9) | 0.0357 (8) | 0.0208 (7) | 0.0091 (7) | 0.0129 (7) |
| C4A | 0.0470 (9) | 0.0488 (10) | 0.0327 (8) | 0.0233 (7) | 0.0079 (7) | 0.0120 (7) |
| C5A | 0.0541 (10) | 0.0457 (9) | 0.0336 (8) | 0.0187 (8) | 0.0067 (7) | 0.0083 (7) |
| C6A | 0.0431 (9) | 0.0471 (9) | 0.0347 (8) | 0.0212 (7) | 0.0078 (7) | 0.0098 (7) |
| C7A | 0.0784 (14) | 0.0613 (12) | 0.0342 (9) | 0.0244 (10) | 0.0027 (9) | 0.0087 (8) |
| C8A | 0.1068 (19) | 0.0696 (15) | 0.0454 (12) | 0.0187 (13) | 0.0015 (12) | 0.0000 (10) |
| C9A | 0.0876 (16) | 0.0564 (12) | 0.0553 (13) | 0.0227 (11) | 0.0051 (11) | −0.0037 (10) |
| C10A | 0.0661 (12) | 0.0478 (10) | 0.0473 (11) | 0.0165 (9) | 0.0033 (9) | 0.0070 (8) |
| C11A | 0.0594 (11) | 0.0505 (11) | 0.0369 (9) | 0.0235 (9) | 0.0089 (8) | 0.0123 (8) |
| C12A | 0.0495 (9) | 0.0511 (10) | 0.0332 (8) | 0.0223 (8) | 0.0074 (7) | 0.0125 (7) |
| C13A | 0.0551 (10) | 0.0524 (10) | 0.0387 (9) | 0.0269 (8) | 0.0093 (7) | 0.0127 (7) |
| C14A | 0.0521 (10) | 0.0617 (11) | 0.0439 (10) | 0.0255 (9) | 0.0056 (8) | 0.0209 (8) |
| C15A | 0.0463 (9) | 0.0604 (11) | 0.0348 (9) | 0.0128 (8) | 0.0048 (7) | 0.0156 (8) |
| C16A | 0.0670 (12) | 0.0566 (11) | 0.0365 (9) | 0.0276 (9) | 0.0089 (8) | 0.0071 (8) |
| C17A | 0.0678 (12) | 0.0525 (11) | 0.0395 (9) | 0.0307 (9) | 0.0093 (8) | 0.0132 (8) |
| C18A | 0.0405 (8) | 0.0478 (9) | 0.0372 (8) | 0.0188 (7) | 0.0087 (6) | 0.0130 (7) |
| C19A | 0.0589 (11) | 0.0514 (10) | 0.0406 (9) | 0.0216 (8) | 0.0104 (8) | 0.0122 (8) |
| C20A | 0.0615 (12) | 0.0474 (11) | 0.0588 (12) | 0.0169 (9) | 0.0015 (9) | 0.0125 (9) |
| C21A | 0.0473 (10) | 0.0568 (11) | 0.0631 (12) | 0.0154 (8) | 0.0053 (8) | 0.0287 (9) |
| C22A | 0.0615 (12) | 0.0668 (13) | 0.0479 (11) | 0.0204 (10) | 0.0159 (9) | 0.0241 (9) |
| C23A | 0.0591 (11) | 0.0542 (11) | 0.0390 (9) | 0.0209 (9) | 0.0129 (8) | 0.0127 (8) |
| C24A | 0.107 (2) | 0.0993 (19) | 0.0368 (11) | 0.0250 (15) | 0.0107 (11) | 0.0079 (11) |
| C25A | 0.0753 (15) | 0.1039 (18) | 0.0522 (12) | 0.0211 (13) | −0.0041 (10) | 0.0379 (12) |
| Cl1B | 0.1053 (5) | 0.0866 (4) | 0.1052 (5) | 0.0675 (4) | 0.0291 (4) | 0.0182 (4) |
| N1B | 0.0523 (8) | 0.0503 (8) | 0.0377 (8) | 0.0228 (7) | 0.0050 (6) | 0.0110 (6) |
| N2B | 0.0718 (11) | 0.0664 (10) | 0.0370 (8) | 0.0402 (8) | 0.0014 (7) | 0.0093 (7) |
| N3B | 0.143 (2) | 0.1261 (18) | 0.0459 (11) | 0.1012 (16) | −0.0007 (11) | 0.0010 (11) |
| N4B | 0.0941 (14) | 0.0619 (11) | 0.0394 (9) | 0.0204 (9) | −0.0015 (8) | 0.0024 (8) |
| C2B | 0.0495 (10) | 0.0521 (10) | 0.0382 (9) | 0.0221 (8) | 0.0057 (7) | 0.0120 (7) |
| C3B | 0.0550 (10) | 0.0518 (10) | 0.0384 (9) | 0.0247 (8) | 0.0066 (7) | 0.0104 (7) |
| C4B | 0.0491 (9) | 0.0439 (9) | 0.0361 (9) | 0.0141 (7) | 0.0070 (7) | 0.0106 (7) |
| C5B | 0.0495 (10) | 0.0476 (10) | 0.0389 (9) | 0.0172 (8) | 0.0032 (7) | 0.0143 (7) |
| C6B | 0.0439 (9) | 0.0432 (9) | 0.0373 (8) | 0.0141 (7) | 0.0074 (7) | 0.0125 (7) |
| C7B | 0.0894 (16) | 0.0775 (14) | 0.0392 (10) | 0.0460 (12) | −0.0002 (10) | 0.0078 (9) |
| C8B | 0.141 (3) | 0.126 (2) | 0.0493 (14) | 0.085 (2) | −0.0014 (14) | 0.0223 (14) |
| C9B | 0.220 (4) | 0.150 (3) | 0.0569 (16) | 0.141 (3) | 0.0197 (19) | 0.0344 (17) |
| C10B | 0.0957 (16) | 0.0833 (15) | 0.0473 (11) | 0.0586 (13) | 0.0085 (11) | 0.0176 (10) |
| C11B | 0.0863 (15) | 0.0743 (14) | 0.0378 (10) | 0.0470 (12) | 0.0013 (9) | 0.0065 (9) |
| C12B | 0.0500 (9) | 0.0447 (9) | 0.0377 (9) | 0.0163 (7) | 0.0062 (7) | 0.0105 (7) |
| C13B | 0.0646 (11) | 0.0480 (10) | 0.0424 (10) | 0.0172 (9) | 0.0101 (8) | 0.0166 (8) |
| C14B | 0.0625 (12) | 0.0432 (10) | 0.0489 (11) | 0.0132 (8) | 0.0051 (8) | 0.0098 (8) |
| C15B | 0.0531 (10) | 0.0502 (10) | 0.0384 (9) | 0.0186 (8) | 0.0019 (7) | 0.0073 (7) |
| C16B | 0.0706 (12) | 0.0524 (11) | 0.0378 (9) | 0.0213 (9) | 0.0080 (8) | 0.0165 (8) |
| C17B | 0.0623 (11) | 0.0414 (9) | 0.0419 (10) | 0.0159 (8) | 0.0070 (8) | 0.0119 (7) |
| C18B | 0.0397 (8) | 0.0428 (9) | 0.0405 (9) | 0.0129 (7) | 0.0071 (7) | 0.0142 (7) |
| C19B | 0.0611 (11) | 0.0651 (12) | 0.0427 (10) | 0.0294 (9) | 0.0107 (8) | 0.0211 (8) |
| C20B | 0.0656 (13) | 0.0754 (14) | 0.0619 (13) | 0.0426 (11) | 0.0125 (10) | 0.0298 (10) |
| C21B | 0.0558 (11) | 0.0543 (11) | 0.0672 (13) | 0.0291 (9) | 0.0181 (9) | 0.0157 (9) |
| C22B | 0.0667 (12) | 0.0579 (12) | 0.0493 (11) | 0.0274 (10) | 0.0166 (9) | 0.0131 (9) |
| C23B | 0.0604 (11) | 0.0508 (10) | 0.0412 (9) | 0.0238 (8) | 0.0089 (8) | 0.0151 (8) |
| C24B | 0.0931 (17) | 0.0963 (18) | 0.0416 (11) | 0.0446 (14) | 0.0099 (11) | 0.0139 (11) |
| C25B | 0.0989 (18) | 0.0630 (14) | 0.0626 (15) | 0.0223 (13) | −0.0107 (13) | −0.0108 (11) |
Geometric parameters (Å, º) top
| Cl1A—C21A | 1.741 (2) | Cl1B—C21B | 1.7444 (19) |
| N1A—C6A | 1.341 (2) | N1B—C6B | 1.342 (2) |
| N1A—C2A | 1.343 (2) | N1B—C2B | 1.345 (2) |
| N2A—C2A | 1.355 (2) | N2B—C2B | 1.354 (2) |
| N2A—C10A | 1.454 (2) | N2B—C10B | 1.455 (2) |
| N2A—C7A | 1.470 (2) | N2B—C7B | 1.476 (2) |
| N3A—C11A | 1.147 (2) | N3B—C11B | 1.140 (3) |
| N4A—C15A | 1.375 (2) | N4B—C15B | 1.386 (2) |
| N4A—C24A | 1.445 (3) | N4B—C25B | 1.436 (3) |
| N4A—C25A | 1.447 (3) | N4B—C24B | 1.439 (3) |
| C2A—C3A | 1.433 (2) | C2B—C3B | 1.432 (2) |
| C3A—C4A | 1.411 (2) | C3B—C4B | 1.409 (2) |
| C3A—C11A | 1.430 (2) | C3B—C11B | 1.436 (3) |
| C4A—C5A | 1.394 (2) | C4B—C5B | 1.392 (2) |
| C4A—C12A | 1.483 (2) | C4B—C12B | 1.481 (2) |
| C5A—C6A | 1.396 (2) | C5B—C6B | 1.392 (2) |
| C5A—H5A | 0.93 | C5B—H5B | 0.93 |
| C6A—C18A | 1.483 (2) | C6B—C18B | 1.488 (2) |
| C7A—C8A | 1.502 (3) | C7B—C8B | 1.511 (3) |
| C7A—H7A | 0.97 | C7B—H7C | 0.97 |
| C7A—H7B | 0.97 | C7B—H7D | 0.97 |
| C8A—C9A | 1.505 (3) | C8B—C9B | 1.416 (4) |
| C8A—H8A | 0.97 | C8B—H8C | 0.97 |
| C8A—H8B | 0.97 | C8B—H8D | 0.97 |
| C9A—C10A | 1.516 (3) | C9B—C10B | 1.516 (3) |
| C9A—H9A | 0.97 | C9B—H9C | 0.97 |
| C9A—H9B | 0.97 | C9B—H9D | 0.97 |
| C10A—H10A | 0.97 | C10B—H10C | 0.97 |
| C10A—H10B | 0.97 | C10B—H10D | 0.97 |
| C12A—C17A | 1.394 (2) | C12B—C13B | 1.387 (2) |
| C12A—C13A | 1.397 (2) | C12B—C17B | 1.398 (2) |
| C13A—C14A | 1.375 (2) | C13B—C14B | 1.377 (3) |
| C13A—H13A | 0.93 | C13B—H13B | 0.93 |
| C14A—C15A | 1.403 (3) | C14B—C15B | 1.393 (3) |
| C14A—H14A | 0.93 | C14B—H14B | 0.93 |
| C15A—C16A | 1.406 (3) | C15B—C16B | 1.403 (3) |
| C16A—C17A | 1.381 (2) | C16B—C17B | 1.376 (2) |
| C16A—H16A | 0.93 | C16B—H16B | 0.93 |
| C17A—H17A | 0.93 | C17B—H17B | 0.93 |
| C18A—C19A | 1.386 (2) | C18B—C19B | 1.388 (2) |
| C18A—C23A | 1.396 (2) | C18B—C23B | 1.391 (2) |
| C19A—C20A | 1.388 (3) | C19B—C20B | 1.383 (3) |
| C19A—H19A | 0.93 | C19B—H19B | 0.93 |
| C20A—C21A | 1.375 (3) | C20B—C21B | 1.374 (3) |
| C20A—H20A | 0.93 | C20B—H20B | 0.93 |
| C21A—C22A | 1.378 (3) | C21B—C22B | 1.371 (3) |
| C22A—C23A | 1.380 (3) | C22B—C23B | 1.383 (2) |
| C22A—H22A | 0.93 | C22B—H22B | 0.93 |
| C23A—H23A | 0.93 | C23B—H23B | 0.93 |
| C24A—H24A | 0.96 | C24B—H24D | 0.96 |
| C24A—H24B | 0.96 | C24B—H24E | 0.96 |
| C24A—H24C | 0.96 | C24B—H24F | 0.96 |
| C25A—H25A | 0.96 | C25B—H25D | 0.96 |
| C25A—H25B | 0.96 | C25B—H25E | 0.96 |
| C25A—H25C | 0.96 | C25B—H25F | 0.96 |
| | | |
| C6A—N1A—C2A | 120.03 (14) | C6B—N1B—C2B | 119.91 (14) |
| C2A—N2A—C10A | 127.52 (15) | C2B—N2B—C10B | 128.03 (16) |
| C2A—N2A—C7A | 119.42 (15) | C2B—N2B—C7B | 119.39 (15) |
| C10A—N2A—C7A | 112.01 (14) | C10B—N2B—C7B | 111.36 (15) |
| C15A—N4A—C24A | 120.53 (19) | C15B—N4B—C25B | 120.12 (18) |
| C15A—N4A—C25A | 120.97 (18) | C15B—N4B—C24B | 121.23 (18) |
| C24A—N4A—C25A | 117.89 (18) | C25B—N4B—C24B | 117.40 (19) |
| N1A—C2A—N2A | 114.81 (14) | N1B—C2B—N2B | 114.66 (15) |
| N1A—C2A—C3A | 121.00 (14) | N1B—C2B—C3B | 120.71 (15) |
| N2A—C2A—C3A | 124.19 (16) | N2B—C2B—C3B | 124.62 (16) |
| C4A—C3A—C11A | 119.50 (14) | C4B—C3B—C2B | 118.92 (15) |
| C4A—C3A—C2A | 118.41 (15) | C4B—C3B—C11B | 118.55 (16) |
| C11A—C3A—C2A | 121.74 (15) | C2B—C3B—C11B | 122.26 (16) |
| C5A—C4A—C3A | 118.23 (14) | C5B—C4B—C3B | 118.28 (15) |
| C5A—C4A—C12A | 119.65 (15) | C5B—C4B—C12B | 119.68 (15) |
| C3A—C4A—C12A | 122.10 (15) | C3B—C4B—C12B | 121.99 (15) |
| C4A—C5A—C6A | 119.77 (15) | C6B—C5B—C4B | 119.50 (16) |
| C4A—C5A—H5A | 120.1 | C6B—C5B—H5B | 120.3 |
| C6A—C5A—H5A | 120.1 | C4B—C5B—H5B | 120.3 |
| N1A—C6A—C5A | 122.09 (16) | N1B—C6B—C5B | 122.60 (15) |
| N1A—C6A—C18A | 114.99 (14) | N1B—C6B—C18B | 115.00 (14) |
| C5A—C6A—C18A | 122.89 (15) | C5B—C6B—C18B | 122.37 (15) |
| N2A—C7A—C8A | 104.21 (17) | N2B—C7B—C8B | 103.24 (17) |
| N2A—C7A—H7A | 110.9 | N2B—C7B—H7C | 111.1 |
| C8A—C7A—H7A | 110.9 | C8B—C7B—H7C | 111.1 |
| N2A—C7A—H7B | 110.9 | N2B—C7B—H7D | 111.1 |
| C8A—C7A—H7B | 110.9 | C8B—C7B—H7D | 111.1 |
| H7A—C7A—H7B | 108.9 | H7C—C7B—H7D | 109.1 |
| C7A—C8A—C9A | 105.59 (18) | C9B—C8B—C7B | 106.3 (2) |
| C7A—C8A—H8A | 110.6 | C9B—C8B—H8C | 110.5 |
| C9A—C8A—H8A | 110.6 | C7B—C8B—H8C | 110.5 |
| C7A—C8A—H8B | 110.6 | C9B—C8B—H8D | 110.5 |
| C9A—C8A—H8B | 110.6 | C7B—C8B—H8D | 110.5 |
| H8A—C8A—H8B | 108.8 | H8C—C8B—H8D | 108.7 |
| C8A—C9A—C10A | 104.69 (17) | C8B—C9B—C10B | 107.5 (2) |
| C8A—C9A—H9A | 110.8 | C8B—C9B—H9C | 110.2 |
| C10A—C9A—H9A | 110.8 | C10B—C9B—H9C | 110.2 |
| C8A—C9A—H9B | 110.8 | C8B—C9B—H9D | 110.2 |
| C10A—C9A—H9B | 110.8 | C10B—C9B—H9D | 110.2 |
| H9A—C9A—H9B | 108.9 | H9C—C9B—H9D | 108.5 |
| N2A—C10A—C9A | 104.13 (16) | N2B—C10B—C9B | 102.85 (17) |
| N2A—C10A—H10A | 110.9 | N2B—C10B—H10C | 111.2 |
| C9A—C10A—H10A | 110.9 | C9B—C10B—H10C | 111.2 |
| N2A—C10A—H10B | 110.9 | N2B—C10B—H10D | 111.2 |
| C9A—C10A—H10B | 110.9 | C9B—C10B—H10D | 111.2 |
| H10A—C10A—H10B | 108.9 | H10C—C10B—H10D | 109.1 |
| N3A—C11A—C3A | 176.9 (2) | N3B—C11B—C3B | 176.5 (3) |
| C17A—C12A—C13A | 116.61 (15) | C13B—C12B—C17B | 116.67 (16) |
| C17A—C12A—C4A | 121.11 (15) | C13B—C12B—C4B | 122.05 (15) |
| C13A—C12A—C4A | 122.12 (15) | C17B—C12B—C4B | 121.18 (15) |
| C14A—C13A—C12A | 121.95 (16) | C14B—C13B—C12B | 122.01 (17) |
| C14A—C13A—H13A | 119.0 | C14B—C13B—H13B | 119.0 |
| C12A—C13A—H13A | 119.0 | C12B—C13B—H13B | 119.0 |
| C13A—C14A—C15A | 121.57 (17) | C13B—C14B—C15B | 121.53 (17) |
| C13A—C14A—H14A | 119.2 | C13B—C14B—H14B | 119.2 |
| C15A—C14A—H14A | 119.2 | C15B—C14B—H14B | 119.2 |
| N4A—C15A—C14A | 121.53 (18) | N4B—C15B—C14B | 121.75 (17) |
| N4A—C15A—C16A | 121.83 (17) | N4B—C15B—C16B | 121.56 (17) |
| C14A—C15A—C16A | 116.62 (16) | C14B—C15B—C16B | 116.69 (16) |
| C17A—C16A—C15A | 121.15 (16) | C17B—C16B—C15B | 121.35 (17) |
| C17A—C16A—H16A | 119.4 | C17B—C16B—H16B | 119.3 |
| C15A—C16A—H16A | 119.4 | C15B—C16B—H16B | 119.3 |
| C16A—C17A—C12A | 122.06 (17) | C16B—C17B—C12B | 121.71 (17) |
| C16A—C17A—H17A | 119.0 | C16B—C17B—H17B | 119.1 |
| C12A—C17A—H17A | 119.0 | C12B—C17B—H17B | 119.1 |
| C19A—C18A—C23A | 118.05 (16) | C19B—C18B—C23B | 118.05 (16) |
| C19A—C18A—C6A | 122.17 (15) | C19B—C18B—C6B | 122.85 (15) |
| C23A—C18A—C6A | 119.77 (15) | C23B—C18B—C6B | 119.10 (15) |
| C18A—C19A—C20A | 121.30 (17) | C20B—C19B—C18B | 120.82 (18) |
| C18A—C19A—H19A | 119.3 | C20B—C19B—H19B | 119.6 |
| C20A—C19A—H19A | 119.3 | C18B—C19B—H19B | 119.6 |
| C21A—C20A—C19A | 119.02 (18) | C21B—C20B—C19B | 119.47 (18) |
| C21A—C20A—H20A | 120.5 | C21B—C20B—H20B | 120.3 |
| C19A—C20A—H20A | 120.5 | C19B—C20B—H20B | 120.3 |
| C20A—C21A—C22A | 121.25 (18) | C22B—C21B—C20B | 121.33 (17) |
| C20A—C21A—Cl1A | 119.70 (16) | C22B—C21B—Cl1B | 118.91 (16) |
| C22A—C21A—Cl1A | 119.05 (16) | C20B—C21B—Cl1B | 119.75 (16) |
| C21A—C22A—C23A | 119.14 (18) | C21B—C22B—C23B | 118.78 (18) |
| C21A—C22A—H22A | 120.4 | C21B—C22B—H22B | 120.6 |
| C23A—C22A—H22A | 120.4 | C23B—C22B—H22B | 120.6 |
| C22A—C23A—C18A | 121.24 (17) | C22B—C23B—C18B | 121.50 (17) |
| C22A—C23A—H23A | 119.4 | C22B—C23B—H23B | 119.2 |
| C18A—C23A—H23A | 119.4 | C18B—C23B—H23B | 119.2 |
| N4A—C24A—H24A | 109.5 | N4B—C24B—H24D | 109.5 |
| N4A—C24A—H24B | 109.5 | N4B—C24B—H24E | 109.5 |
| H24A—C24A—H24B | 109.5 | H24D—C24B—H24E | 109.5 |
| N4A—C24A—H24C | 109.5 | N4B—C24B—H24F | 109.5 |
| H24A—C24A—H24C | 109.5 | H24D—C24B—H24F | 109.5 |
| H24B—C24A—H24C | 109.5 | H24E—C24B—H24F | 109.5 |
| N4A—C25A—H25A | 109.5 | N4B—C25B—H25D | 109.5 |
| N4A—C25A—H25B | 109.5 | N4B—C25B—H25E | 109.5 |
| H25A—C25A—H25B | 109.5 | H25D—C25B—H25E | 109.5 |
| N4A—C25A—H25C | 109.5 | N4B—C25B—H25F | 109.5 |
| H25A—C25A—H25C | 109.5 | H25D—C25B—H25F | 109.5 |
| H25B—C25A—H25C | 109.5 | H25E—C25B—H25F | 109.5 |
| | | |
| C6A—N1A—C2A—N2A | 175.55 (15) | C6B—N1B—C2B—N2B | 179.49 (16) |
| C6A—N1A—C2A—C3A | −4.6 (2) | C6B—N1B—C2B—C3B | −0.9 (3) |
| C10A—N2A—C2A—N1A | −168.95 (17) | C10B—N2B—C2B—N1B | −169.62 (19) |
| C7A—N2A—C2A—N1A | −1.7 (2) | C7B—N2B—C2B—N1B | −3.4 (3) |
| C10A—N2A—C2A—C3A | 11.2 (3) | C10B—N2B—C2B—C3B | 10.8 (3) |
| C7A—N2A—C2A—C3A | 178.52 (16) | C7B—N2B—C2B—C3B | 177.07 (18) |
| N1A—C2A—C3A—C4A | 8.1 (2) | N1B—C2B—C3B—C4B | 2.1 (3) |
| N2A—C2A—C3A—C4A | −172.10 (16) | N2B—C2B—C3B—C4B | −178.35 (18) |
| N1A—C2A—C3A—C11A | −165.16 (15) | N1B—C2B—C3B—C11B | −171.72 (19) |
| N2A—C2A—C3A—C11A | 14.7 (3) | N2B—C2B—C3B—C11B | 7.8 (3) |
| C11A—C3A—C4A—C5A | 167.86 (16) | C2B—C3B—C4B—C5B | −3.1 (3) |
| C2A—C3A—C4A—C5A | −5.5 (2) | C11B—C3B—C4B—C5B | 170.93 (18) |
| C11A—C3A—C4A—C12A | −13.7 (2) | C2B—C3B—C4B—C12B | 174.50 (15) |
| C2A—C3A—C4A—C12A | 172.88 (15) | C11B—C3B—C4B—C12B | −11.4 (3) |
| C3A—C4A—C5A—C6A | −0.1 (2) | C3B—C4B—C5B—C6B | 3.1 (3) |
| C12A—C4A—C5A—C6A | −178.52 (15) | C12B—C4B—C5B—C6B | −174.62 (15) |
| C2A—N1A—C6A—C5A | −1.3 (2) | C2B—N1B—C6B—C5B | 0.9 (3) |
| C2A—N1A—C6A—C18A | −179.35 (14) | C2B—N1B—C6B—C18B | −177.10 (15) |
| C4A—C5A—C6A—N1A | 3.7 (2) | C4B—C5B—C6B—N1B | −2.0 (3) |
| C4A—C5A—C6A—C18A | −178.42 (15) | C4B—C5B—C6B—C18B | 175.85 (15) |
| C2A—N2A—C7A—C8A | −176.48 (18) | C2B—N2B—C7B—C8B | −176.8 (2) |
| C10A—N2A—C7A—C8A | −7.3 (2) | C10B—N2B—C7B—C8B | −8.4 (3) |
| N2A—C7A—C8A—C9A | 23.7 (2) | N2B—C7B—C8B—C9B | 24.0 (3) |
| C7A—C8A—C9A—C10A | −31.2 (3) | C7B—C8B—C9B—C10B | −30.8 (4) |
| C2A—N2A—C10A—C9A | 156.24 (18) | C2B—N2B—C10B—C9B | 158.1 (2) |
| C7A—N2A—C10A—C9A | −11.8 (2) | C7B—N2B—C10B—C9B | −9.1 (3) |
| C8A—C9A—C10A—N2A | 26.2 (2) | C8B—C9B—C10B—N2B | 24.7 (4) |
| C5A—C4A—C12A—C17A | −42.0 (2) | C5B—C4B—C12B—C13B | 128.23 (19) |
| C3A—C4A—C12A—C17A | 139.55 (18) | C3B—C4B—C12B—C13B | −49.4 (3) |
| C5A—C4A—C12A—C13A | 133.25 (18) | C5B—C4B—C12B—C17B | −47.9 (2) |
| C3A—C4A—C12A—C13A | −45.2 (2) | C3B—C4B—C12B—C17B | 134.50 (19) |
| C17A—C12A—C13A—C14A | 0.9 (3) | C17B—C12B—C13B—C14B | −0.3 (3) |
| C4A—C12A—C13A—C14A | −174.58 (16) | C4B—C12B—C13B—C14B | −176.56 (17) |
| C12A—C13A—C14A—C15A | −1.4 (3) | C12B—C13B—C14B—C15B | −1.7 (3) |
| C24A—N4A—C15A—C14A | −179.0 (2) | C25B—N4B—C15B—C14B | −2.5 (3) |
| C25A—N4A—C15A—C14A | −8.1 (3) | C24B—N4B—C15B—C14B | 164.4 (2) |
| C24A—N4A—C15A—C16A | 2.6 (3) | C25B—N4B—C15B—C16B | 176.8 (2) |
| C25A—N4A—C15A—C16A | 173.49 (19) | C24B—N4B—C15B—C16B | −16.4 (3) |
| C13A—C14A—C15A—N4A | −178.23 (17) | C13B—C14B—C15B—N4B | −178.31 (19) |
| C13A—C14A—C15A—C16A | 0.3 (3) | C13B—C14B—C15B—C16B | 2.4 (3) |
| N4A—C15A—C16A—C17A | 179.74 (17) | N4B—C15B—C16B—C17B | 179.46 (19) |
| C14A—C15A—C16A—C17A | 1.3 (3) | C14B—C15B—C16B—C17B | −1.2 (3) |
| C15A—C16A—C17A—C12A | −1.7 (3) | C15B—C16B—C17B—C12B | −0.7 (3) |
| C13A—C12A—C17A—C16A | 0.6 (3) | C13B—C12B—C17B—C16B | 1.4 (3) |
| C4A—C12A—C17A—C16A | 176.17 (17) | C4B—C12B—C17B—C16B | 177.76 (17) |
| N1A—C6A—C18A—C19A | −167.70 (15) | N1B—C6B—C18B—C19B | −166.42 (17) |
| C5A—C6A—C18A—C19A | 14.3 (2) | C5B—C6B—C18B—C19B | 15.6 (3) |
| N1A—C6A—C18A—C23A | 12.1 (2) | N1B—C6B—C18B—C23B | 12.9 (2) |
| C5A—C6A—C18A—C23A | −165.89 (16) | C5B—C6B—C18B—C23B | −165.10 (16) |
| C23A—C18A—C19A—C20A | −0.1 (3) | C23B—C18B—C19B—C20B | −2.5 (3) |
| C6A—C18A—C19A—C20A | 179.66 (16) | C6B—C18B—C19B—C20B | 176.84 (17) |
| C18A—C19A—C20A—C21A | 0.5 (3) | C18B—C19B—C20B—C21B | 1.4 (3) |
| C19A—C20A—C21A—C22A | −0.7 (3) | C19B—C20B—C21B—C22B | 0.2 (3) |
| C19A—C20A—C21A—Cl1A | 179.11 (15) | C19B—C20B—C21B—Cl1B | −178.85 (17) |
| C20A—C21A—C22A—C23A | 0.4 (3) | C20B—C21B—C22B—C23B | −0.7 (3) |
| Cl1A—C21A—C22A—C23A | −179.40 (15) | Cl1B—C21B—C22B—C23B | 178.37 (15) |
| C21A—C22A—C23A—C18A | 0.0 (3) | C21B—C22B—C23B—C18B | −0.4 (3) |
| C19A—C18A—C23A—C22A | −0.2 (3) | C19B—C18B—C23B—C22B | 2.0 (3) |
| C6A—C18A—C23A—C22A | −179.97 (16) | C6B—C18B—C23B—C22B | −177.35 (17) |
Hydrogen-bond geometry (Å, º) top
| D—H···A | D—H | H···A | D···A | D—H···A |
| C19B—H19B···N3A | 0.93 | 2.55 | 3.461 (3) | 167 |
| C19A—H19A···N3Bi | 0.93 | 2.47 | 3.378 (3) | 164 |
| C24B—H24F···N3Aii | 0.96 | 2.59 | 3.457 (3) | 150 |
| C23A—H23A···N1A | 0.93 | 2.42 | 2.755 (2) | 101 |
| C23B—H23B···N1B | 0.93 | 2.41 | 2.745 (3) | 101 |
| Symmetry codes: (i) x, y−1, z; (ii) −x+1, −y+1, −z+1. |
Experimental details
| Crystal data |
| Chemical formula | C24H23ClN4 |
| Mr | 402.91 |
| Crystal system, space group | Triclinic, P1 |
| Temperature (K) | 293 |
| a, b, c (Å) | 11.1490 (1), 13.7330 (1), 14.6666 (1) |
| α, β, γ (°) | 100.487 (1), 93.240 (1), 107.014 (1) |
| V (Å3) | 2096.93 (2) |
| Z | 4 |
| Radiation type | Mo Kα |
| µ (mm−1) | 0.20 |
| Crystal size (mm) | 0.48 × 0.44 × 0.26 |
| |
| Data collection |
| Diffractometer | Siemens SMART CCD area-detector
diffractometer |
| Absorption correction | – |
No. of measured, independent and
observed [I > 2σ(I)] reflections | 14581, 9579, 6307 |
| Rint | 0.048 |
| (sin θ/λ)max (Å−1) | 0.667 |
| |
| Refinement |
| R[F2 > 2σ(F2)], wR(F2), S | 0.054, 0.160, 0.97 |
| No. of reflections | 9579 |
| No. of parameters | 527 |
| H-atom treatment | H-atom parameters constrained |
| Δρmax, Δρmin (e Å−3) | 0.35, −0.32 |
Selected geometric parameters (Å, º) top| N2A—C2A | 1.355 (2) | N2B—C7B | 1.476 (2) |
| N2A—C10A | 1.454 (2) | N4B—C15B | 1.386 (2) |
| N2A—C7A | 1.470 (2) | N4B—C25B | 1.436 (3) |
| N4A—C15A | 1.375 (2) | N4B—C24B | 1.439 (3) |
| N4A—C24A | 1.445 (3) | C7B—C8B | 1.511 (3) |
| N4A—C25A | 1.447 (3) | C8B—C9B | 1.416 (4) |
| N2B—C2B | 1.354 (2) | C9B—C10B | 1.516 (3) |
| N2B—C10B | 1.455 (2) | | |
| | | |
| N1A—C2A—N2A | 114.81 (14) | N1B—C2B—N2B | 114.66 (15) |
| N2A—C2A—C3A | 124.19 (16) | N2B—C2B—C3B | 124.62 (16) |
| N1A—C6A—C18A | 114.99 (14) | N1B—C6B—C18B | 115.00 (14) |
| C5A—C6A—C18A | 122.89 (15) | C5B—C6B—C18B | 122.37 (15) |
Hydrogen-bond geometry (Å, º) top
| D—H···A | D—H | H···A | D···A | D—H···A |
| C19B—H19B···N3A | 0.93 | 2.55 | 3.461 (3) | 167 |
| C19A—H19A···N3Bi | 0.93 | 2.47 | 3.378 (3) | 164 |
| C24B—H24F···N3Aii | 0.96 | 2.59 | 3.457 (3) | 150 |
| C23A—H23A···N1A | 0.93 | 2.42 | 2.755 (2) | 101 |
| C23B—H23B···N1B | 0.93 | 2.41 | 2.745 (3) | 101 |
| Symmetry codes: (i) x, y−1, z; (ii) −x+1, −y+1, −z+1. |
 N hydrogen bonds, through the formation of hydrogen-bonded molecular chains.
N hydrogen bonds, through the formation of hydrogen-bonded molecular chains.![[sigma]](//journals.iucr.org/logos/entities/sigma_rmgif.gif) (C-C) = 0.003 Å
(C-C) = 0.003 ÅAlert Level C:
![[Figure 1]](http://journals.iucr.org/e/issues/2002/12/00/ci6179/ci6179fig1thm.gif)

 journal menu
journal menu














Nicotine-containing compounds show interesting biological properties. The nicotinic acid derivative N,N-diethylnicotinamide, which is commonly known as DENA, has the respiratory stimulating property (Hokelek & Necefouglu, 1996). Niacin is a vitamin that contains nicotinamide, deficiency of which makes the body to lose copper thereby creating the pellagra disease (Hokelek & Necefouglu, 1999). The redox pair NAD+ (nicotinamide adenine dinucleotide) molecule, which is the oxidized form of the coenzyme NADH, plays a key role in energy-producing processes (Stryer, 1988) and in the redox processes catalysed by various protein families, the dehydrogenases being the largest group (Guillot et al., 2000). The X-ray investigation of the title compound, (I), was taken up as a part of our study on the structures of a nicotine derivatives,
The asymmetric unit of the title compound, (I), contains two molecules; the corresponding bond lengths and angles of these two molecules agree with each other, except for some deviations in Csp3—Csp3 distances of the pyrrolidine ring. A molecular fit for these two molecules shows a weighted r.m.s. deviation of 0.52 Å. The geometry of the two molecules slightly differ in the orientation of the dimethylaminophenyl ring and the nitrile group with respect to the pyridine ring. The N2—C2—C3 angles [124.2 (2) and 124.6 (2)°] are wider than the N1—C2—N2 angles [114.8 (1) and 114.7 (2)°], as a result of the repulsive force exerted by the nitrile group on the pyrrolidine ring. Also, as a result of the steric repulsion between atoms H5 and H19 (H5···H19 = 2.18 Å each), the exocyclic angles C5—C6—C18 [122.9 (2) and 122.4 (2)°] are wider compared with N1—C6—C18 [115.0 (1) and 115.0 (1)°]. Similar distortions in the bond angles have been reported previously for a related structure (Thinagar et al., 2002). The pyrrolidine rings of both the molecules adopt half-chair conformations, with ΔC2(N2) values of 0.012 (1) and 0.002 (1) (Nardelli, 1983), and mean planes through them form dihedral angles of 18.4 (1) and 13.1 (1)° with the pyridine rings. The twisting of the chlorophenyl ring from the pyridine ring is less [12.83 (9) and 14.36 (9)°] compared to that of the other phenyl ring [43.14 (9) and 49.36 (9)°].
In the asymmetric unit, the two crystallographically independent molecules are linked by C19B—H19B···N3A hydrogen bonds. In the crystal, these pairs translated a unit along the b axis are linked by C19A—H19A···N3B hydrogen bonds to form an infinite one-dimensional molecular chain (Table 2). The molecules of these chains are interlinked to those in the adjacent inversion-related chain by C24B—H24F···N3A(1 − x, 1 − y, 1 − z) hydrogen bonds to form double-chain structures.