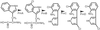issue contents
August 2004 issue

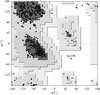
Cover illustration: Sharply kinked DNA decamers form virtual double helices, with Sac7d molecules bound to alternate sides of the DNA (p. 1381).
research papers
The aldose reductase–IDD552 complex confirms the possibility of direct proton donation from the active-site residue Tyr48.
Open  access
access
 access
accessThe small-molecule topology generator PRODRG is described, which takes input from existing coordinates or various two-dimensional formats and automatically generates coordinates and molecular topologies suitable for X-ray refinement of protein–ligand complexes.
A single-crystal neutron diffraction study has been carried out at 1.6 Å resolution on a mutant rubredoxin from the hyperthermophilic archeon P. furiosus.
The structure of E. faecalis haem catalase was determined by molecular replacement and refined to 2.3 Å resolution using synchrotron data.
PDB reference: catalase, 1si8, r1si8sf
The crystal structures of two Sac7d–DNA decamer complexes were determined in two different unit cells. The DNA decamers are more bent than the octamer, and they form triple base pairs in the lattice contact regions.
Structures of active-site mutants have definitively established the identity and roles of the substrate-binding groups in aspartate-β-semialdehyde dehydrogenase.
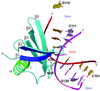
The structure of the CH domain of Rng2 is described at 2.21 Å resolution. Two features that are new to CH domains are revealed: a carboxyl-terminal tail that wraps around the putative actin-binding site ABS1 and ABS2 from dystrophin, and two loop conformations between Tyr92 and Arg99. This may have useful implications in determining the actin-binding orientation from single CH domains.
Two crystal structures of a jacalin-related lectin from A. hirsuta, which is specific for galactose, in complex with methyl-α-D-galactose are reported. The lectin crystallized in two orthorhombic forms and one hexagonal form under similar conditions. The structure was solved using the molecular-replacement method using the jacalin structure as a search model and the two orthorhombic forms were refined using data to 2.5 and 3.0 Å resolution, respectively.
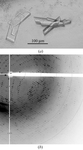
A simple and fast desiccation procedure was successfully applied to three cases of poorly diffracting protein crystals leading to a dramatic improvement in their diffraction. This could become a last-resort protocol to be systematically applied to non-diffracting crystals.
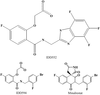
Crystal structure of binary complex of calf spleen purine-nucleoside phosphorylase with a potent multisubstrate analogue inhibitor 9-(5,5-difluoro-5-phosphonopentyl)guanine has been refined to 2.2 Å resolution. The present structure may be used in the rational, structure-based design of new PNP inhibitors with potential medical applications.
PDB reference: PNP–DFPP-G complex, 1v48, r1v48sf
crystallization papers
This report details the production, purification, crystallization and preliminary diffraction analysis of CD3∊γ and CD3∊δ complexed with OKT3.
The membrane-bound DegS protease from E. coli has been crystallized in a free and a substrate-bound form. The structure has been solved by molecular replacement and the solution was verified by radiation-induced phasing (RIP).
E. coli ketol-acid reductoisomerase has been crystallized under a wide range of conditions and in several crystal forms. Most of the crystals diffract poorly but we have collected several data sets at better than 3.2 Å resolution.
Methylmalonate-semialdehyde dehydrogenase which is a singular member of the aldehyde dehydrogenase family was overexpressed and crystallized. Crystals diffract to 2.5 Å and belong to the orthorhombic space group P212121.
Open  access
access
 access
accessCrystallization of PrnA (tryptophan 7-halogenase), an enzyme that regioselectively chlorinates tryptophan, cocrystallized with tryptophan and FAD is reported.
Crystallization of the yeast S. cerevisiae mitochondrial F1-ATPase with improvements leading to diffraction from 3.3 to 2.8 Å.
The crystallization and preliminary X-ray analysis of carboxypeptidase 1 from T. thermophilus have been conducted.
UDP-glucose pyrophosphorylase from H. pylori has been overexpressed, purified and crystallized. The crystals were analyzed by X-ray crystallography to a resolution of 2.9 Å.
The glycine-cleavage system component T-protein from P. horikoshii OT3 was overexpressed and purified. Orthorhombic crystals were obtained and native data were collected to 1.5 Å resolution using synchrotron radiation.
Phenoxazinone synthase was crystallized using the sitting-drop vapour-diffusion method. Crystals were formed under two separate conditions, with crystals from each condition assigned to a different space group. Diffraction data from the two different crystals forms were measured to resolution limits of 3.35 and 2.30 Å and processed in space groups R32 and P1, respectively.
The active oxidized form of nitroalkane oxidase, an FAD-dependent enzyme isolated from Fusarium oxysporum, has been crystallized in three space groups. A multiwavelength data set from Se-Met enriched enzyme is suitable for determining the structure.
This study describes the first crystallization and preliminary X-ray analysis of a family 52 glycoside hydrolase, the β-D-xylosidase from G. stearothermophilus T-6. Purified native enzyme was crystallized and belongs to space group P1. The native crystal diffracted X-rays to a resolution of 2.0 Å.
A dimeric double-headed Kunitz-type potato serine protease inhibitor isolated from potato tubers was crystallized. Preliminary X-ray data were collected to 1.8 Å. The crystals are monoclinic, space group P21, with unit-cell parameters a = 54.82, b = 93.92, c = 55.44 Å, β = 100.7°.
The crystallization and X-ray data analysis of crystals of the outer membrane pyoverdine receptor FpvA from P. aeruginosa are described. Crystals of the in vivo FpvA–pyoverdine complex and the selenomethionine-labelled protein were obtained in the presence of C8E5 detergent. Both crystal forms belong to space group C2.
The catalytic module of β-1,3-xylanase from Vibrio sp. AX-4 has been crystallized and X-ray diffraction data have been collected to 1.44 Å.
Open  access
access
 access
accessCrystals of MHC H-2Kd complexed with an HBV immunodominant nonapeptide were produced that diffracted X-rays to 2.06 Å.
The DNA-repair enzyme endonuclease VIII (Nei) from E. coli has been crystallized in its DNA-free form. The new I222 crystals enabled a full X-ray diffraction data measurement for wild-type Nei at 2.6 Å resolution, for the Nei-E2A mutant at 2.25 Å resolution and for the Nei-R252A mutant at 2.05 Å resolution.
Crystals of an enzyme from the crenarchaeon P. aerophilum that catalyzes both phosphoglucose isomerase and phosphomannose isomerase activities have been obtained. They belong to space group P21 and diffract to at least 1.6 Å resolution.
Archaeal ATP synthase catalytic subunit A from P. horikoshii OT3 was overexpressed, purified and crystallized. X-ray diffraction data were collected to 2.55 Å resolution.
A novel hyperthermophilic archaeal Rieske protein, sulredoxin from S. tokodaii, has been overproduced, purified and crystallized in the oxidized conformation. The cubic crystals diffract to beyond 2.0 Å resolution using synchrotron radiation.
Open  access
access
 access
accessAn alkaline mannanase from a Bacillus sp. was crystallized and the crystals diffracted to 1.65 Å using synchrotron radiation. Phasing was successfully carried out using the molecular-replacement method.
The seed lectin of C. gladiata was crystallized in orthorhombic space group C2221. X-ray data were collected at 2.3 Å resolution.
Glutathione S-transferase from the pathogenic human filarial parasite O. volvulus has been crystallized, allowing its X-ray structure determination.
Monodehydroascorbate (MDA) radical reductase from cucumber has been crystallized and X-ray diffraction data have been collected to 2.4 Å.
A 1.95 Å resolution data set was collected from a crystal of a recombinant Z. mays cytokinin oxidase (ZmCKO1). This enzyme degrades cytokinins, hormones that are involved in plant growth and development.
short communications
Open  access
access
 access
accessThe structure of a tetragonal crystal form of C. botulinum C3 exoenzyme has been determined. NAD-induced main-chain flexure is concentrated in two lobes peripheral to a rigid β-stranded core.
PDB reference: C3 exoenzyme, tetragonal form, 1uzi, r1uzisf
Excellent gadolinium-derivative crystals of hen egg-white lysozyme were obtained by co-crystallizing the protein with 100 mM Gd-HPDO3A using the lipidic cubic phase method known to be effective for crystallizing membrane proteins.


 journal menu
journal menu