issue contents
January 2016 issue

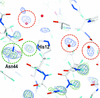
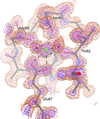
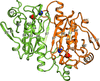
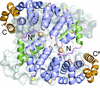
Cover illustration: Surface representation of the molecular packing in the nucleoprotein C-terminal domain of the Bundibugyo species of Ebolavirus (BDBV NPCt) looking down the sixfold screw axis visualizing the solvent channels. This protein is critical for the assembly and replication of the virus and consequently is recognized as a suitable drug target. Importantly, the Ebolavirus NPCt has also been identified as a possible target for the development of species-specific diagnostic tests and the structural characterization of NPCt from different Ebolavirus species is relevant to this potential application (Baker et al., p. 40).
editorial
Free 

The Acta Cryst. D Editors look forward to a change in scope and a new subtitle for the journal.
research papers
Open  access
access
 access
accessA high-density sample mount has been developed for efficient goniometer-based sample delivery at synchrotron and XFEL sources.
Five crystal structures of D-alanine-D-alanine ligase from Y. pestis have been determined at 1.7–2.5 Å resolution: apo, AMP-bound, ADP-bound, adenosine 5′-(β,γ-imido)triphosphate-bound, and D-alanyl-D-alanine- and ADP-bound structures. The serine loop is mainly responsible for the conformational change in the substrate nucleotide phosphates.
This article presents the crystal structure of full-length VgrG1 from P. aeruginosa, a component of the type VI secretion-system machinery essential in bacterial infection.
PDB reference: VgrG1, 4uhv
Open  access
access
 access
accessA specialized filter for finding lattices in images of three-dimensional nanocrystals devoid of any contrast is described.
The structure of the extramembrane portion of the CcdA-family member Rv2874 from M. tuberculosis reveals a previously unseen juxtaposition of a thioredoxin-like domain with a carbohydrate-binding module, suggesting a role in cell-wall carbohydrate processing.
Crystal structures of the C-terminal domains of the Ebolavirus nucleoproteins (NPCt) from the Bundibugyo and Taï Forest species (BDBV and TAFV, respectively) have been determined. The structures show high similarity to that reported for the Zaire Ebolavirus NPCt. However, NMR data revealed that the corresponding domain from the NP of the related MARV species of Marburgvirus is distinctly different.
Wild-type and mutant structures of B. licheniformis trehalose-6-phosphate hydrolase suggest new insight into the characteristics and substrate specificity of trehalose-6-phosphate hydrolase.
The deuterium/hydrogen neutron contrast map, which was calculated using subtraction in real space, demonstrated the powerful detectability of H/D exchange in the neutron structure determination of a ribonuclease A crystal.
Experimental phase information at 1.0 Å resolution was used to validate a more complete multiple-conformation model of disorder in Ca2+-calmodulin and identified bound 2-methyl-2,4-pentanediol (MPD) in multiple regions of the protein. This bound MPD is likely to contribute to some of the previously noted peculiar features of the Ca2+-calmodulin crystal structure.
Download citation


Download citation


Open  access
access
 access
accessA method for performing high-throughput in situ serial X-ray crystallography with soluble and membrane proteins in the lipid cubic phase at cryogenic temperatures (100 K) is described. It works with nanogram to single-digit microgram quantities of protein and lipid (and ligand when present), and is compatible with both high-resolution native data collection and experimental phasing without the need for crystal harvesting.
Open  access
access
 access
accessCrystal structures of LukE and LukD from S. aureus are reported at 3.20 and 1.70 Å resolution, respectively.
The structural implications of incorporating spectroscopic reporter unnatural amino acids into proteins is explored using X-ray crystallography. Four protein crystal structures with 4-cyano-L-phenylalanine or 4-ethynyl-L-phenylalanine incorporated into green fluorescent protein at two unique sites are described.
Open  access
access
 access
accessThe structure of the spliceosomal U4 snRNP core domain has been re-refined following molecular replacement using the minimal U1 snRNP as a search model. RNA-omit maps show that the U4 Sm-site nucleotides AAUUUUU are bound to the seven Sm proteins SmF–SmE–SmG–SmD3–SmB–SmD1–SmD2 in the same manner as the U1 Sm-site nucleotides AAUUUGU except at the U-to-G substitution in SmD1.
A procedure is suggested for ab initio phasing of diffraction data for single-particle studies. The procedure is based on a Monte Carlo-type search for connected binary masks approximating electron-density distribution.
The crystal structure of TSA2 reveals a novel conformation of a strictly conserved proline and explains the differences in pKa between yeast peroxiredoxins.
PDB reference: S. cerevisiae TSA2, 5dvb
Open  access
access
 access
accessDesigned armadillo-repeat proteins are promising scaffolds for modular peptide-recognition systems. The crystal structures of His-YIIIM4AII, His-YIIIM5AII and YIIIM5AII highlight structural heterogeneity in full-consensus designs and aid the improvement of future constructs.
scientific comment
Open  access
access
 access
accessThe default geometry restraints used in Phenix for the protein backbone have been upgraded to account for the known conformation-dependencies of bond angles and lengths.
addenda and errata
Free 

book reviews
Free 



 journal menu
journal menu