issue contents
March 2018 issue
From crystal to structure with CCP4 (Part 2)
Proceedings of the CCP4 Study Weekend edited by Charles Ballard, Mike Hough and Keith Wilson
research papers
Open  access
access
 access
accessNew developments in the MrBUMP pipeline, including graphical interaction with search models using CCP4mg and the use of ensemble search models, are reported.
Open  access
access
 access
accessNovel ways to produce search models from distant homologues for molecular replacement are presented.
Open  access
access
 access
accessThe ARCIMBOLDO method of phasing through the location of small fragments combined with density modification and autotracing is particularly suited to helical structures, but coiled coils remain challenging. Features designed for solving coiled coils at resolutions of up to 3 Å were tested on a pool of 150 structures.
Open  access
access
 access
accessA new pipeline to solve structures by molecular replacement with ideal protein fragments is described and benchmarked against two test sets of mixed α/β and all-β folds at relatively high resolution.
Open  access
access
 access
accessHere, a macromolecule-centred approach to three-dimensional structure determination as implemented in REFMAC5 is considered. The use of restraints to transfer chemical and structural information during macromolecular refinement, and how different sources of information can be combined in order to achieve models that are more consistent with data derived from a variety of experimental techniques, including macromolecular crystallography, cryo-EM and NMR spectroscopy, are discussed.
Open  access
access
 access
accessBetter metrics are required to be able to assess small-molecule ligands in macromolecular structures in Worldwide Protein Data Bank validation reports. The local ligand density fit (LLDF) score currently used to assess ligand electron-density fit outliers produces a substantial number of false positives and false negatives.
Open  access
access
 access
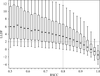
accessThe use of validation metrics to rank macromolecular structures, as well as a web tool to investigate trends in and correlations between different properties and validation metrics, are described.


 journal menu
journal menu