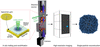issue contents
June 2023 issue

Cover illustration: The Collaborative Computational Project No. 4 (CCP4) suite is a collection of macromolecular crystallography software brought together by familiar execution routines, a set of common libraries, and graphical interfaces. A new article by Agirre et al. [(2023), Acta Cryst. D79, 449–461], which is intended as a general literature citation for the use of the CCP4 software suite in structure determination, gives the reader a general overview of the programs in the suite.
scientific commentaries
Open  access
access
 access
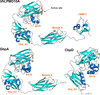
accessA new chitin-active AA10 lytic polysaccharide monooxygenase from the marine bacterium Vibrio campbellii is described in the paper by Zhou et al. [(2023), Acta Cryst. D79, 479–497].
Open  access
access
 access
accessThe evolution of the Collaborative Computational Project No. 4 (CCP4) has been described in a new article by Agirre et al. [(2023). Acta Cryst. D79, 449–461] that should provide the definitive reference for the CCP4 suite of programs.
CCP4
Open  access
access
 access
accessThis article describes the Collaborative Computational Project No. 4 (CCP4). It is intended as a general literature citation for the use of the CCP4 software suite in structure determination.
Open  access
access
 access
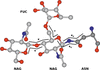
accessThe Privateer software allows structural biologists to evaluate and improve the atomic structures of carbohydrates, including N-glycans. This software has recently been extended to check glycan composition through the use of glycomics data, and the broadening of its scope is presented in this article.
CCP-EM
Open  access
access
 access
accessNear-atomic resolution reconstructions can be obtained from in situ melted and revitrified cryo samples. Revitrification results in a more homogeneous angular distribution.
research papers
Chitin-active lytic polysaccharide monooxygenase (AA10 LPMO) is one of the key enzymes involved in the recycling of chitin biomass. A new marine chitin-active AA10 LPMO from V. campbellii could serve as a powerful biocatalyst in the bioconversion of chitin for future bioenergy production.
Open  access
access
 access
accessThe latch domain in reverse gyrases mediates the cooperation of the helicase and topoisomerase domains. In Thermotoga maritima reverse gyrase this latch can be designed into a minimal β-bulge loop that has natural precedence in Thermosipho africanus reverse gyrase. The properties and functionalities of different latch domains across reverse gyrases are discussed.
Open  access
access
 access
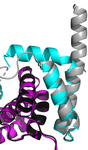
accessSctX proteins are secreted substrates and are essential components of type III secretion systems. The dedicated chaperone SctY escorts SctX proteins to the secretion machinery, where the hydrophobic helix at the C-terminus of SctX mediates recognition by the export apparatus. New crystal structures suggest that in the absence of the export gate this hydrophobic helix kinks and folds back onto the chaperone, presumably to shield it from the solvent.
Open  access
access
 access
accessBiochemical and structural analyses of cysteine synthase, the key enzyme in cysteine biosynthesis, from the protozoan pathogens Trypanosoma cruzi, T. theileri and Leishmania infantum are presented. This enzyme is a potential drug target for neglected tropical diseases such as Chagas disease and leishmaniasis.
Superoxide dismutase 1 (SOD1) is a therapeutic drug target against amyotrophic lateral sclerosis. In silico and in vitro studies indicate binding of paliperidone at Trp32 of SOD1, an important site for oxidative stress-induced aggregation. Significant interactions with SOD1 and low-micromolar binding affinity make paliperidone a potential lead for the development of drug candidates that can prevent SOD1 fibrillation.
PDB reference: human superoxide dismutase, complex with paliperidone, 8gsq
Open  access
access
 access
accessStructural investigation of a presumed fungal glucuronoyl esterase reveals a classical serine hydrolase active site but an unusual ligand-binding site. Functional analysis showed a lack of activity on a wide array of substrates commonly utilized by this family of enzymes. It is hypothesized that this enzyme needs complex plant cell-wall substructures for activity.
PDB reference: LfCE15C, 8b48

 journal menu
journal menu