- editorial
- research papers
- research communications
- research papers
- crystallization communications
- research papers
- crystallization communications
- research papers
- crystallization communications
- research papers
- crystallization communications
- feature articles
- protein structure communications
- crystallization communications
- protein structure communications
- crystallization papers
- structural genomics papers
- abstracts
issue contents
Coronaviruses (2020)
Structural biology has an important role in fighting the current COVID-19 pandemic caused by the coronavirus known as SARS-CoV-2. This virtual issue presents articles and abstracts on coronaviruses that have been published in IUCr journals.
Many structural biology initiatives are under way to combat COVID-19 and here are just a few examples: a giant fragment screening exercise led by researchers at Diamond Light Source (https://covid.postera.ai/covid); the CASP initiative to model SARS-2-CoV targets (http://predictioncenter.org/caspcommons/index.cgi); a protopedia page on COVID-19 [http://proteopedia.org/w/Coronavirus_Disease_2019_(COVID-19)]; a large collaborative effort on rerefined structures with a webpage featuring numerous links (https://github.com/thorn-lab/coronavirus_structural_task_force); the COVID-19 Portal, which will bring together relevant datasets submitted to EMBL-EBI and other major centres for biomedical data (https://www.ebi.ac.uk/covid-19).
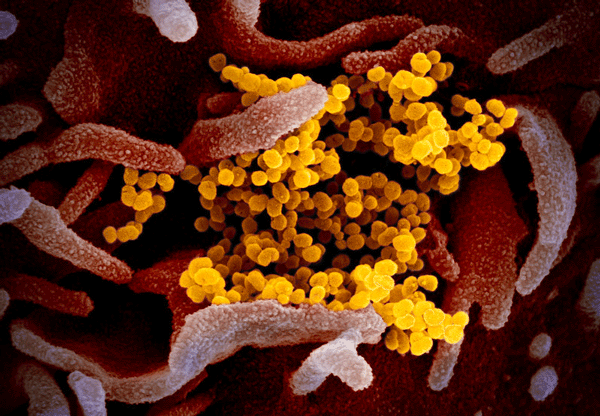
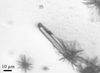
Cover illustration: This scanning electron microscope image shows SARS-CoV-2 (yellow) - the virus that causes COVID-19 - isolated from a patient in the USA, emerging from the surface of cells (pink) cultured in the laboratory. Credit: NIAID-RML
editorial
research articles
 access
access access
access
 access
accessrelated abstracts














 menu
menu