Buy article online - an online subscription or single-article purchase is required to access this article.
The synthesis,
1H and
13C NMR spectra, and X-ray structures are described for three dialkoxy ethynylnitrobenzenes that differ only in the length of the alkoxy chain, namely 1-ethynyl-2-nitro-4,5-dipropoxybenzene, C
14H
17NO
4, 1,2-dibutoxy-4-ethynyl-5-nitrobenzene, C
16H
21NO
4, and 1-ethynyl-2-nitro-4,5-dipentoxybenzene, C
18H
25NO
4. Despite the subtle changes in molecular structure, the crystal structures of the three compounds display great diversity. Thus, 1-ethynyl-2-nitro-4,5-dipropoxybenzene crystallizes in the trigonal crystal system in the space group
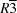
, with
Z = 18, 1,2-dibutoxy-4-ethynyl-5-nitrobenzene crystallizes in the monoclinic crystal system in the space group
P2
1/
c, with
Z = 4, and 1-ethynyl-2-nitro-4,5-dipentoxybenzene crystallizes in the triclinic crystal system in the space group
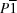
, with
Z = 2. The crystal structure of 1-ethynyl-2-nitro-4,5-dipropoxybenzene is dominated by planar hexamers formed by a bifurcated alkoxy
sp-C—H

O,O′ interaction, while the structure of the dibutoxy analogue is dominated by planar ribbons of molecules linked by a similar bifurcated alkoxy
sp-C—H

O,O′ interaction. In contrast, the dipentoxy analogue forms ribbons of molecules alternately connected by a self-complementary
sp-C—H

O
2N interaction and a self-complementary
sp2-C—H

O
2N interaction. Disordered solvent was included in the crystals of 1-ethynyl-2-nitro-4,5-dipropoxybenzene and its contribution was removed during refinement.
Supporting information
CCDC references: 1572979; 1572978; 1572977
For all structures, data collection: Bruker SMART; cell refinement: Bruker SMART; data reduction: Bruker SAINT; program(s) used to solve structure: SHELXT 2014/5 (Sheldrick, 2015); program(s) used to refine structure: SHELXL2017/1 (Sheldrick, 2015a); molecular graphics: X-SEED (Barbour, 2001); software used to prepare material for publication: X-SEED (Barbour, 2001).
Crystal data top
| C14H17NO4 | Dx = 1.234 Mg m−3 |
| Mr = 263.29 | Mo Kα radiation, λ = 0.71073 Å |
| Trigonal, R3:H | Cell parameters from 5437 reflections |
| a = 26.6774 (16) Å | θ = 2.6–27.1° |
| c = 10.3451 (6) Å | µ = 0.09 mm−1 |
| V = 6376.1 (8) Å3 | T = 100 K |
| Z = 18 | Cut rectangular prism, colourless |
| F(000) = 2520 | 0.07 × 0.07 × 0.04 mm |
Data collection top
Bruker APEXII CCD area-detector
diffractometer | 3172 independent reflections |
| Radiation source: fine-focus sealed tube | 2420 reflections with I > 2σ(I) |
| Graphite monochromator | Rint = 0.039 |
| Detector resolution: 8.3660 pixels mm-1 | θmax = 27.2°, θmin = 1.5° |
| φ and ω scans | h = −34→34 |
Absorption correction: multi-scan
(SADABS Version 2014; Bruker AXS) | k = −34→34 |
| Tmin = 0.697, Tmax = 0.746 | l = −13→13 |
| 28334 measured reflections | |
Refinement top
| Refinement on F2 | 2 restraints |
| Least-squares matrix: full | Hydrogen site location: mixed |
| R[F2 > 2σ(F2)] = 0.047 | H atoms treated by a mixture of independent and constrained refinement |
| wR(F2) = 0.138 | w = 1/[σ2(Fo2) + (0.0842P)2 + 1.9773P]
where P = (Fo2 + 2Fc2)/3 |
| S = 1.06 | (Δ/σ)max = 0.001 |
| 3172 reflections | Δρmax = 0.43 e Å−3 |
| 180 parameters | Δρmin = −0.20 e Å−3 |
Special details top
Geometry. All esds (except the esd in the dihedral angle between two l.s. planes)
are estimated using the full covariance matrix. The cell esds are taken
into account individually in the estimation of esds in distances, angles
and torsion angles; correlations between esds in cell parameters are only
used when they are defined by crystal symmetry. An approximate (isotropic)
treatment of cell esds is used for estimating esds involving l.s. planes. |
Refinement. # SQUEEZE RESULTS
# Note: Data are Listed for all Voids in the P1 Unit Cell
# i.e. Centre of Gravity, Solvent Accessible Volume,
# Recovered number of Electrons in the Void and
# Details about the Squeezed Material
loop_
_platon_squeeze_void_nr
_platon_squeeze_void_average_x
_platon_squeeze_void_average_y
_platon_squeeze_void_average_z
_platon_squeeze_void_volume
_platon_squeeze_void_count_electrons
_platon_squeeze_void_content
1 0.000 0.000 0.000 125 38 ' '
2 0.333 0.667 0.667 115 38 ' '
3 0.667 0.333 0.333 115 38 ' ' _platon_squeeze_details
'Reasonable that the squeezed solvent corresponds to dichloromethane (42 e)' |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2) top| | x | y | z | Uiso*/Ueq | |
| O1 | 0.80137 (4) | 0.82653 (4) | 0.29721 (12) | 0.0429 (3) | |
| N1 | 0.75096 (5) | 0.80819 (5) | 0.32978 (11) | 0.0256 (3) | |
| C1 | 0.73231 (5) | 0.70715 (5) | 0.33025 (12) | 0.0206 (3) | |
| O2 | 0.73202 (4) | 0.83983 (4) | 0.36150 (11) | 0.0375 (3) | |
| C2 | 0.71127 (5) | 0.74567 (5) | 0.32948 (12) | 0.0212 (3) | |
| O3 | 0.55388 (4) | 0.64527 (4) | 0.32599 (9) | 0.0261 (2) | |
| C3 | 0.65221 (6) | 0.72722 (5) | 0.32879 (12) | 0.0212 (3) | |
| O4 | 0.59014 (4) | 0.57189 (4) | 0.32933 (9) | 0.0244 (2) | |
| C4 | 0.61230 (5) | 0.66862 (5) | 0.32747 (12) | 0.0209 (3) | |
| C5 | 0.63200 (5) | 0.62827 (5) | 0.32965 (12) | 0.0209 (3) | |
| C6 | 0.69061 (5) | 0.64767 (5) | 0.33122 (12) | 0.0205 (3) | |
| H6 | 0.703270 | 0.620132 | 0.333023 | 0.025* | |
| C7 | 0.79185 (6) | 0.72157 (5) | 0.33451 (12) | 0.0232 (3) | |
| C8 | 0.83787 (6) | 0.72495 (6) | 0.33970 (14) | 0.0286 (3) | |
| C9 | 0.53135 (6) | 0.68428 (6) | 0.31049 (13) | 0.0252 (3) | |
| H9A | 0.543558 | 0.704456 | 0.226250 | 0.030* | |
| H9B | 0.546036 | 0.713660 | 0.380149 | 0.030* | |
| C10 | 0.46645 (6) | 0.64833 (6) | 0.31691 (16) | 0.0336 (3) | |
| H10A | 0.454772 | 0.629926 | 0.403095 | 0.040* | |
| H10B | 0.452595 | 0.617229 | 0.251226 | 0.040* | |
| C11 | 0.43856 (7) | 0.68527 (7) | 0.2930 (2) | 0.0480 (5) | |
| H11A | 0.453678 | 0.717186 | 0.355410 | 0.072* | |
| H11B | 0.396514 | 0.661425 | 0.303249 | 0.072* | |
| H11C | 0.447534 | 0.700984 | 0.205091 | 0.072* | |
| C12 | 0.60887 (6) | 0.52969 (5) | 0.33039 (13) | 0.0239 (3) | |
| H12A | 0.630358 | 0.533250 | 0.411084 | 0.029* | |
| H12B | 0.634935 | 0.536310 | 0.256252 | 0.029* | |
| C13 | 0.55624 (6) | 0.47007 (6) | 0.32152 (15) | 0.0290 (3) | |
| H13A | 0.534786 | 0.466784 | 0.240927 | 0.035* | |
| H13B | 0.530215 | 0.463725 | 0.395589 | 0.035* | |
| C14 | 0.57451 (7) | 0.42451 (6) | 0.32240 (17) | 0.0374 (4) | |
| H14A | 0.598579 | 0.429645 | 0.246451 | 0.056* | |
| H14B | 0.540037 | 0.385883 | 0.320216 | 0.056* | |
| H14C | 0.596699 | 0.428606 | 0.401088 | 0.056* | |
| H3 | 0.6402 (8) | 0.7547 (7) | 0.3249 (16) | 0.045* | |
| H8 | 0.8740 (5) | 0.7274 (8) | 0.3423 (16) | 0.045* | |
Atomic displacement parameters (Å2) top| | U11 | U22 | U33 | U12 | U13 | U23 |
| O1 | 0.0265 (6) | 0.0225 (5) | 0.0748 (8) | 0.0085 (4) | 0.0085 (5) | 0.0029 (5) |
| N1 | 0.0259 (6) | 0.0183 (6) | 0.0312 (6) | 0.0101 (5) | −0.0031 (5) | −0.0002 (4) |
| C1 | 0.0218 (6) | 0.0205 (6) | 0.0188 (6) | 0.0101 (5) | −0.0019 (5) | −0.0009 (5) |
| O2 | 0.0337 (6) | 0.0217 (5) | 0.0597 (7) | 0.0158 (4) | −0.0025 (5) | −0.0055 (5) |
| C2 | 0.0239 (6) | 0.0166 (6) | 0.0209 (6) | 0.0085 (5) | −0.0011 (5) | 0.0002 (5) |
| O3 | 0.0192 (5) | 0.0203 (5) | 0.0393 (6) | 0.0102 (4) | −0.0003 (4) | 0.0022 (4) |
| C3 | 0.0245 (6) | 0.0200 (6) | 0.0215 (6) | 0.0128 (5) | −0.0019 (5) | 0.0000 (5) |
| O4 | 0.0202 (5) | 0.0162 (4) | 0.0347 (5) | 0.0075 (4) | −0.0003 (4) | 0.0005 (4) |
| C4 | 0.0190 (6) | 0.0221 (6) | 0.0214 (6) | 0.0101 (5) | −0.0005 (5) | 0.0004 (5) |
| C5 | 0.0223 (6) | 0.0186 (6) | 0.0199 (6) | 0.0087 (5) | −0.0014 (5) | −0.0006 (5) |
| C6 | 0.0230 (6) | 0.0177 (6) | 0.0220 (6) | 0.0110 (5) | −0.0011 (5) | −0.0013 (5) |
| C7 | 0.0261 (7) | 0.0166 (6) | 0.0250 (6) | 0.0092 (5) | −0.0019 (5) | −0.0018 (5) |
| C8 | 0.0229 (7) | 0.0235 (7) | 0.0380 (8) | 0.0105 (6) | −0.0026 (6) | −0.0020 (6) |
| C9 | 0.0244 (7) | 0.0240 (6) | 0.0309 (7) | 0.0149 (6) | −0.0005 (5) | 0.0022 (5) |
| C10 | 0.0225 (7) | 0.0295 (7) | 0.0496 (9) | 0.0136 (6) | 0.0007 (6) | 0.0032 (6) |
| C11 | 0.0307 (8) | 0.0336 (8) | 0.0846 (13) | 0.0199 (7) | −0.0147 (8) | −0.0088 (8) |
| C12 | 0.0256 (7) | 0.0188 (6) | 0.0289 (7) | 0.0123 (5) | 0.0001 (5) | −0.0003 (5) |
| C13 | 0.0264 (7) | 0.0204 (7) | 0.0366 (7) | 0.0089 (6) | −0.0018 (6) | −0.0015 (6) |
| C14 | 0.0381 (8) | 0.0195 (7) | 0.0520 (10) | 0.0124 (6) | 0.0014 (7) | 0.0003 (6) |
Geometric parameters (Å, º) top
| O1—N1 | 1.2262 (14) | C9—C10 | 1.5037 (18) |
| N1—O2 | 1.2255 (14) | C9—H9A | 0.9900 |
| N1—C2 | 1.4619 (16) | C9—H9B | 0.9900 |
| C1—C2 | 1.3955 (17) | C10—C11 | 1.523 (2) |
| C1—C6 | 1.4108 (16) | C10—H10A | 0.9900 |
| C1—C7 | 1.4361 (18) | C10—H10B | 0.9900 |
| C2—C3 | 1.3963 (18) | C11—H11A | 0.9800 |
| O3—C4 | 1.3587 (14) | C11—H11B | 0.9800 |
| O3—C9 | 1.4476 (15) | C11—H11C | 0.9800 |
| C3—C4 | 1.3831 (17) | C12—C13 | 1.5087 (17) |
| C3—H3 | 0.936 (14) | C12—H12A | 0.9900 |
| O4—C5 | 1.3527 (15) | C12—H12B | 0.9900 |
| O4—C12 | 1.4422 (14) | C13—C14 | 1.5191 (19) |
| C4—C5 | 1.4148 (17) | C13—H13A | 0.9900 |
| C5—C6 | 1.3796 (17) | C13—H13B | 0.9900 |
| C6—H6 | 0.9500 | C14—H14A | 0.9800 |
| C7—C8 | 1.1862 (19) | C14—H14B | 0.9800 |
| C8—H8 | 0.935 (9) | C14—H14C | 0.9800 |
| | | |
| O2—N1—O1 | 123.12 (11) | C9—C10—C11 | 111.07 (12) |
| O2—N1—C2 | 118.26 (11) | C9—C10—H10A | 109.4 |
| O1—N1—C2 | 118.62 (10) | C11—C10—H10A | 109.4 |
| C2—C1—C6 | 116.55 (11) | C9—C10—H10B | 109.4 |
| C2—C1—C7 | 126.96 (11) | C11—C10—H10B | 109.4 |
| C6—C1—C7 | 116.46 (10) | H10A—C10—H10B | 108.0 |
| C1—C2—C3 | 122.61 (11) | C10—C11—H11A | 109.5 |
| C1—C2—N1 | 120.76 (11) | C10—C11—H11B | 109.5 |
| C3—C2—N1 | 116.62 (11) | H11A—C11—H11B | 109.5 |
| C4—O3—C9 | 117.71 (9) | C10—C11—H11C | 109.5 |
| C4—C3—C2 | 119.58 (11) | H11A—C11—H11C | 109.5 |
| C4—C3—H3 | 120.8 (11) | H11B—C11—H11C | 109.5 |
| C2—C3—H3 | 119.5 (11) | O4—C12—C13 | 108.62 (10) |
| C5—O4—C12 | 116.89 (10) | O4—C12—H12A | 110.0 |
| O3—C4—C3 | 125.21 (11) | C13—C12—H12A | 110.0 |
| O3—C4—C5 | 115.38 (10) | O4—C12—H12B | 110.0 |
| C3—C4—C5 | 119.40 (11) | C13—C12—H12B | 110.0 |
| O4—C5—C6 | 124.60 (11) | H12A—C12—H12B | 108.3 |
| O4—C5—C4 | 115.58 (10) | C12—C13—C14 | 109.95 (12) |
| C6—C5—C4 | 119.82 (11) | C12—C13—H13A | 109.7 |
| C5—C6—C1 | 122.02 (11) | C14—C13—H13A | 109.7 |
| C5—C6—H6 | 119.0 | C12—C13—H13B | 109.7 |
| C1—C6—H6 | 119.0 | C14—C13—H13B | 109.7 |
| C8—C7—C1 | 170.34 (13) | H13A—C13—H13B | 108.2 |
| C7—C8—H8 | 179.0 (11) | C13—C14—H14A | 109.5 |
| O3—C9—C10 | 107.23 (10) | C13—C14—H14B | 109.5 |
| O3—C9—H9A | 110.3 | H14A—C14—H14B | 109.5 |
| C10—C9—H9A | 110.3 | C13—C14—H14C | 109.5 |
| O3—C9—H9B | 110.3 | H14A—C14—H14C | 109.5 |
| C10—C9—H9B | 110.3 | H14B—C14—H14C | 109.5 |
| H9A—C9—H9B | 108.5 | | |
| | | |
| C3—C2—N1—O2 | 17.89 (17) | C1—C2—N1—O1 | 18.77 (18) |
Hydrogen-bond geometry (Å, º) top
| D—H···A | D—H | H···A | D···A | D—H···A |
| C8—H8···O3i | 0.94 (1) | 2.48 (1) | 3.4001 (16) | 167 (2) |
| C8—H8···O4i | 0.94 (1) | 2.62 (2) | 3.3267 (16) | 133 (1) |
| C3—H3···O2ii | 0.94 (1) | 2.61 (1) | 3.5459 (16) | 177 (2) |
| Symmetry codes: (i) y+1/3, −x+y+2/3, −z+2/3; (ii) −x+4/3, −y+5/3, −z+2/3. |
1,2-dibutoxy-4-ethynyl-5-nitrobenzene (2)
top
Crystal data top
| C16H21NO4 | F(000) = 624 |
| Mr = 291.34 | Dx = 1.273 Mg m−3 |
| Monoclinic, P21/c | Mo Kα radiation, λ = 0.71073 Å |
| a = 8.5209 (5) Å | Cell parameters from 3448 reflections |
| b = 18.0914 (10) Å | θ = 2.3–26.4° |
| c = 9.9579 (5) Å | µ = 0.09 mm−1 |
| β = 97.967 (1)° | T = 100 K |
| V = 1520.24 (14) Å3 | Irregular block, colourless |
| Z = 4 | 0.60 × 0.34 × 0.02 mm |
Data collection top
Bruker APEXII CCD area-detector
diffractometer | 3370 independent reflections |
| Radiation source: fine-focus sealed tube | 2449 reflections with I > 2σ(I) |
| Graphite monochromator | Rint = 0.047 |
| Detector resolution: 8.3660 pixels mm-1 | θmax = 27.2°, θmin = 2.3° |
| φ and ω scans | h = −10→10 |
Absorption correction: multi-scan
(SADABS Version 2014; Bruker AXS) | k = −23→23 |
| Tmin = 0.682, Tmax = 0.746 | l = −12→12 |
| 19579 measured reflections | |
Refinement top
| Refinement on F2 | 2 restraints |
| Least-squares matrix: full | Hydrogen site location: mixed |
| R[F2 > 2σ(F2)] = 0.041 | H atoms treated by a mixture of independent and constrained refinement |
| wR(F2) = 0.103 | w = 1/[σ2(Fo2) + (0.0447P)2 + 0.5063P]
where P = (Fo2 + 2Fc2)/3 |
| S = 1.02 | (Δ/σ)max < 0.001 |
| 3370 reflections | Δρmax = 0.25 e Å−3 |
| 198 parameters | Δρmin = −0.18 e Å−3 |
Special details top
Geometry. All esds (except the esd in the dihedral angle between two l.s. planes)
are estimated using the full covariance matrix. The cell esds are taken
into account individually in the estimation of esds in distances, angles
and torsion angles; correlations between esds in cell parameters are only
used when they are defined by crystal symmetry. An approximate (isotropic)
treatment of cell esds is used for estimating esds involving l.s. planes. |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2) top| | x | y | z | Uiso*/Ueq | |
| O1 | 0.24462 (15) | 0.41330 (6) | −0.09599 (12) | 0.0331 (3) | |
| N1 | 0.22644 (15) | 0.34657 (7) | −0.09358 (12) | 0.0194 (3) | |
| C1 | 0.41750 (17) | 0.33670 (8) | 0.12054 (14) | 0.0153 (3) | |
| O2 | 0.13376 (14) | 0.31327 (7) | −0.17849 (12) | 0.0335 (3) | |
| C2 | 0.31940 (17) | 0.30296 (8) | 0.01366 (14) | 0.0153 (3) | |
| O3 | 0.38628 (12) | 0.10631 (5) | 0.09739 (10) | 0.0170 (2) | |
| C3 | 0.30591 (17) | 0.22633 (8) | 0.00230 (14) | 0.0156 (3) | |
| H3 | 0.2381 (18) | 0.2081 (9) | −0.0705 (14) | 0.019* | |
| H8 | 0.4862 (18) | 0.5273 (7) | 0.1967 (15) | 0.019* | |
| O4 | 0.57880 (12) | 0.16462 (5) | 0.28966 (10) | 0.0169 (2) | |
| C4 | 0.39137 (16) | 0.18124 (8) | 0.09726 (14) | 0.0146 (3) | |
| C5 | 0.49571 (16) | 0.21339 (8) | 0.20434 (14) | 0.0145 (3) | |
| C6 | 0.50568 (17) | 0.28947 (8) | 0.21533 (14) | 0.0150 (3) | |
| H6 | 0.573938 | 0.310557 | 0.288897 | 0.018* | |
| C7 | 0.43705 (17) | 0.41478 (8) | 0.14269 (14) | 0.0183 (3) | |
| C8 | 0.46506 (19) | 0.47713 (9) | 0.17401 (16) | 0.0227 (3) | |
| C9 | 0.27503 (17) | 0.07150 (8) | −0.00558 (14) | 0.0168 (3) | |
| H9A | 0.307039 | 0.079990 | −0.096176 | 0.020* | |
| H9B | 0.167922 | 0.092592 | −0.005009 | 0.020* | |
| C10 | 0.27354 (17) | −0.01003 (8) | 0.02436 (14) | 0.0167 (3) | |
| H10A | 0.236740 | −0.017855 | 0.113319 | 0.020* | |
| H10B | 0.382713 | −0.029677 | 0.030183 | 0.020* | |
| C11 | 0.16582 (18) | −0.05231 (8) | −0.08441 (15) | 0.0190 (3) | |
| H11A | 0.057299 | −0.031722 | −0.092033 | 0.023* | |
| H11B | 0.204412 | −0.045622 | −0.172931 | 0.023* | |
| C12 | 0.16069 (19) | −0.13435 (8) | −0.05226 (16) | 0.0231 (4) | |
| H12A | 0.266801 | −0.155571 | −0.050285 | 0.035* | |
| H12B | 0.086938 | −0.159234 | −0.122124 | 0.035* | |
| H12C | 0.125019 | −0.141126 | 0.036353 | 0.035* | |
| C13 | 0.69850 (17) | 0.19390 (8) | 0.39333 (14) | 0.0161 (3) | |
| H13A | 0.648403 | 0.224452 | 0.458108 | 0.019* | |
| H13B | 0.773813 | 0.225262 | 0.351626 | 0.019* | |
| C14 | 0.78460 (17) | 0.12937 (8) | 0.46602 (14) | 0.0175 (3) | |
| H14A | 0.844815 | 0.102735 | 0.402880 | 0.021* | |
| H14B | 0.706485 | 0.094605 | 0.495658 | 0.021* | |
| C15 | 0.89834 (18) | 0.15516 (9) | 0.58933 (15) | 0.0200 (3) | |
| H15A | 0.974564 | 0.190909 | 0.559722 | 0.024* | |
| H15B | 0.837535 | 0.180877 | 0.653162 | 0.024* | |
| C16 | 0.9892 (2) | 0.09090 (10) | 0.66219 (17) | 0.0288 (4) | |
| H16A | 0.914086 | 0.055400 | 0.691711 | 0.043* | |
| H16B | 1.059132 | 0.109511 | 0.741391 | 0.043* | |
| H16C | 1.052701 | 0.066468 | 0.600332 | 0.043* | |
Atomic displacement parameters (Å2) top| | U11 | U22 | U33 | U12 | U13 | U23 |
| O1 | 0.0427 (8) | 0.0206 (7) | 0.0324 (7) | 0.0030 (5) | −0.0069 (6) | 0.0077 (5) |
| N1 | 0.0194 (7) | 0.0200 (7) | 0.0184 (7) | 0.0039 (5) | 0.0015 (5) | 0.0017 (5) |
| C1 | 0.0153 (7) | 0.0148 (7) | 0.0164 (7) | −0.0001 (6) | 0.0039 (6) | 0.0014 (6) |
| O2 | 0.0371 (7) | 0.0301 (7) | 0.0271 (6) | 0.0032 (5) | −0.0167 (5) | −0.0001 (5) |
| C2 | 0.0147 (7) | 0.0178 (8) | 0.0136 (7) | 0.0037 (6) | 0.0021 (6) | 0.0032 (6) |
| O3 | 0.0199 (5) | 0.0123 (5) | 0.0169 (5) | −0.0012 (4) | −0.0043 (4) | −0.0008 (4) |
| C3 | 0.0132 (7) | 0.0193 (8) | 0.0137 (7) | −0.0004 (6) | 0.0002 (6) | −0.0016 (6) |
| O4 | 0.0198 (5) | 0.0130 (5) | 0.0155 (5) | −0.0010 (4) | −0.0060 (4) | 0.0011 (4) |
| C4 | 0.0137 (7) | 0.0159 (7) | 0.0145 (7) | 0.0001 (6) | 0.0022 (5) | −0.0002 (6) |
| C5 | 0.0139 (7) | 0.0163 (7) | 0.0128 (7) | 0.0015 (6) | 0.0006 (5) | 0.0021 (6) |
| C6 | 0.0138 (7) | 0.0181 (8) | 0.0130 (7) | −0.0011 (6) | 0.0008 (5) | −0.0010 (6) |
| C7 | 0.0179 (8) | 0.0208 (8) | 0.0156 (7) | 0.0029 (6) | 0.0006 (6) | 0.0035 (6) |
| C8 | 0.0261 (9) | 0.0172 (8) | 0.0235 (8) | 0.0015 (7) | −0.0016 (6) | 0.0026 (7) |
| C9 | 0.0152 (7) | 0.0184 (8) | 0.0152 (7) | −0.0011 (6) | −0.0032 (6) | −0.0015 (6) |
| C10 | 0.0152 (7) | 0.0165 (8) | 0.0172 (7) | −0.0002 (6) | −0.0018 (6) | −0.0001 (6) |
| C11 | 0.0207 (8) | 0.0163 (8) | 0.0186 (7) | 0.0000 (6) | −0.0022 (6) | −0.0018 (6) |
| C12 | 0.0237 (8) | 0.0184 (8) | 0.0257 (8) | −0.0008 (6) | −0.0026 (7) | −0.0012 (7) |
| C13 | 0.0167 (7) | 0.0161 (7) | 0.0143 (7) | −0.0031 (6) | −0.0025 (6) | −0.0010 (6) |
| C14 | 0.0190 (8) | 0.0166 (8) | 0.0156 (7) | 0.0005 (6) | −0.0022 (6) | 0.0006 (6) |
| C15 | 0.0183 (7) | 0.0249 (9) | 0.0154 (7) | 0.0026 (6) | −0.0029 (6) | −0.0007 (6) |
| C16 | 0.0256 (9) | 0.0363 (10) | 0.0220 (8) | 0.0039 (7) | −0.0056 (7) | 0.0075 (7) |
Geometric parameters (Å, º) top
| O1—N1 | 1.2178 (16) | C10—C11 | 1.5248 (19) |
| N1—O2 | 1.2314 (16) | C10—H10A | 0.9900 |
| N1—C2 | 1.4683 (18) | C10—H10B | 0.9900 |
| C1—C2 | 1.399 (2) | C11—C12 | 1.520 (2) |
| C1—C6 | 1.4103 (19) | C11—H11A | 0.9900 |
| C1—C7 | 1.436 (2) | C11—H11B | 0.9900 |
| C2—C3 | 1.394 (2) | C12—H12A | 0.9800 |
| O3—C4 | 1.3564 (17) | C12—H12B | 0.9800 |
| O3—C9 | 1.4412 (16) | C12—H12C | 0.9800 |
| C3—C4 | 1.378 (2) | C13—C14 | 1.509 (2) |
| C3—H3 | 0.923 (13) | C13—H13A | 0.9900 |
| O4—C5 | 1.3546 (16) | C13—H13B | 0.9900 |
| O4—C13 | 1.4473 (16) | C14—C15 | 1.5281 (19) |
| C4—C5 | 1.4152 (19) | C14—H14A | 0.9900 |
| C5—C6 | 1.382 (2) | C14—H14B | 0.9900 |
| C6—H6 | 0.9500 | C15—C16 | 1.524 (2) |
| C7—C8 | 1.186 (2) | C15—H15A | 0.9900 |
| C8—H8 | 0.947 (13) | C15—H15B | 0.9900 |
| C9—C10 | 1.505 (2) | C16—H16A | 0.9800 |
| C9—H9A | 0.9900 | C16—H16B | 0.9800 |
| C9—H9B | 0.9900 | C16—H16C | 0.9800 |
| | | |
| O1—N1—O2 | 122.87 (13) | C12—C11—C10 | 111.85 (12) |
| O1—N1—C2 | 119.33 (12) | C12—C11—H11A | 109.2 |
| O2—N1—C2 | 117.80 (12) | C10—C11—H11A | 109.2 |
| C2—C1—C6 | 116.84 (13) | C12—C11—H11B | 109.2 |
| C2—C1—C7 | 126.19 (13) | C10—C11—H11B | 109.2 |
| C6—C1—C7 | 116.97 (13) | H11A—C11—H11B | 107.9 |
| C3—C2—C1 | 121.96 (13) | C11—C12—H12A | 109.5 |
| C3—C2—N1 | 116.44 (13) | C11—C12—H12B | 109.5 |
| C1—C2—N1 | 121.60 (13) | H12A—C12—H12B | 109.5 |
| C4—O3—C9 | 117.00 (11) | C11—C12—H12C | 109.5 |
| C4—C3—C2 | 120.21 (13) | H12A—C12—H12C | 109.5 |
| C4—C3—H3 | 122.8 (10) | H12B—C12—H12C | 109.5 |
| C2—C3—H3 | 117.0 (10) | O4—C13—C14 | 107.86 (11) |
| C5—O4—C13 | 117.67 (11) | O4—C13—H13A | 110.1 |
| O3—C4—C3 | 125.34 (13) | C14—C13—H13A | 110.1 |
| O3—C4—C5 | 115.27 (12) | O4—C13—H13B | 110.1 |
| C3—C4—C5 | 119.39 (13) | C14—C13—H13B | 110.1 |
| O4—C5—C6 | 125.30 (13) | H13A—C13—H13B | 108.4 |
| O4—C5—C4 | 115.09 (12) | C13—C14—C15 | 111.17 (12) |
| C6—C5—C4 | 119.61 (13) | C13—C14—H14A | 109.4 |
| C5—C6—C1 | 121.94 (13) | C15—C14—H14A | 109.4 |
| C5—C6—H6 | 119.0 | C13—C14—H14B | 109.4 |
| C1—C6—H6 | 119.0 | C15—C14—H14B | 109.4 |
| C8—C7—C1 | 172.37 (16) | H14A—C14—H14B | 108.0 |
| C7—C8—H8 | 178.5 (10) | C16—C15—C14 | 111.92 (13) |
| O3—C9—C10 | 108.06 (11) | C16—C15—H15A | 109.2 |
| O3—C9—H9A | 110.1 | C14—C15—H15A | 109.2 |
| C10—C9—H9A | 110.1 | C16—C15—H15B | 109.2 |
| O3—C9—H9B | 110.1 | C14—C15—H15B | 109.2 |
| C10—C9—H9B | 110.1 | H15A—C15—H15B | 107.9 |
| H9A—C9—H9B | 108.4 | C15—C16—H16A | 109.5 |
| C9—C10—C11 | 111.82 (12) | C15—C16—H16B | 109.5 |
| C9—C10—H10A | 109.3 | H16A—C16—H16B | 109.5 |
| C11—C10—H10A | 109.3 | C15—C16—H16C | 109.5 |
| C9—C10—H10B | 109.3 | H16A—C16—H16C | 109.5 |
| C11—C10—H10B | 109.3 | H16B—C16—H16C | 109.5 |
| H10A—C10—H10B | 107.9 | | |
| | | |
| C1—C2—N1—O1 | 6.1 (2) | C3—C2—N1—O2 | 6.1 (2) |
Hydrogen-bond geometry (Å, º) top
| D—H···A | D—H | H···A | D···A | D—H···A |
| C8—H8···O3i | 0.95 (1) | 2.61 (1) | 3.3835 (19) | 139 (1) |
| C8—H8···O4i | 0.95 (1) | 2.55 (1) | 3.4371 (19) | 156 (1) |
| Symmetry code: (i) −x+1, y+1/2, −z+1/2. |
Crystal data top
| C18H25NO4 | Z = 2 |
| Mr = 319.39 | F(000) = 344 |
| Triclinic, P1 | Dx = 1.181 Mg m−3 |
| a = 4.570 (4) Å | Mo Kα radiation, λ = 0.71073 Å |
| b = 9.415 (8) Å | Cell parameters from 852 reflections |
| c = 20.968 (18) Å | θ = 2.3–25.6° |
| α = 94.003 (12)° | µ = 0.08 mm−1 |
| β = 90.758 (12)° | T = 100 K |
| γ = 93.435 (11)° | Square, colourless |
| V = 898.2 (14) Å3 | 0.46 × 0.14 × 0.14 mm |
Data collection top
Bruker APEXII CCD area-detector
diffractometer | 2914 independent reflections |
| Radiation source: fine-focus sealed tube | 1604 reflections with I > 2σ(I) |
| Graphite monochromator | Rint = 0.055 |
| Detector resolution: 8.3660 pixels mm-1 | θmax = 24.4°, θmin = 2.0° |
| φ and ω scans | h = −5→5 |
Absorption correction: multi-scan
(SADABS Version 2014; Bruker AXS) | k = −10→10 |
| Tmin = 0.382, Tmax = 0.746 | l = −24→23 |
| 5380 measured reflections | |
Refinement top
| Refinement on F2 | 2 restraints |
| Least-squares matrix: full | Hydrogen site location: mixed |
| R[F2 > 2σ(F2)] = 0.085 | H atoms treated by a mixture of independent and constrained refinement |
| wR(F2) = 0.262 | w = 1/[σ2(Fo2) + (0.1254P)2 + 0.8636P]
where P = (Fo2 + 2Fc2)/3 |
| S = 1.03 | (Δ/σ)max < 0.001 |
| 2914 reflections | Δρmax = 0.36 e Å−3 |
| 216 parameters | Δρmin = −0.32 e Å−3 |
Special details top
Geometry. All esds (except the esd in the dihedral angle between two l.s. planes)
are estimated using the full covariance matrix. The cell esds are taken
into account individually in the estimation of esds in distances, angles
and torsion angles; correlations between esds in cell parameters are only
used when they are defined by crystal symmetry. An approximate (isotropic)
treatment of cell esds is used for estimating esds involving l.s. planes. |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2) top| | x | y | z | Uiso*/Ueq | |
| O1 | 0.5970 (9) | 0.2146 (5) | 0.48201 (18) | 0.0588 (11) | |
| N1 | 0.4403 (9) | 0.2986 (5) | 0.5092 (2) | 0.0370 (10) | |
| C1 | 0.6541 (10) | 0.2513 (5) | 0.6159 (2) | 0.0315 (12) | |
| O2 | 0.2526 (9) | 0.3609 (5) | 0.48100 (18) | 0.0590 (12) | |
| C2 | 0.4709 (10) | 0.3307 (5) | 0.5786 (2) | 0.0295 (11) | |
| O3 | 0.1873 (6) | 0.5814 (3) | 0.70199 (15) | 0.0326 (8) | |
| C3 | 0.3139 (10) | 0.4410 (5) | 0.6053 (2) | 0.0298 (11) | |
| O4 | 0.5189 (7) | 0.4440 (3) | 0.77303 (15) | 0.0345 (9) | |
| C4 | 0.3319 (9) | 0.4762 (5) | 0.6707 (2) | 0.0266 (11) | |
| C5 | 0.5148 (9) | 0.4001 (5) | 0.7095 (2) | 0.0290 (11) | |
| C6 | 0.6704 (10) | 0.2917 (5) | 0.6818 (2) | 0.0310 (11) | |
| H6 | 0.793656 | 0.242129 | 0.708311 | 0.037* | |
| C7 | 0.8201 (11) | 0.1340 (5) | 0.5926 (2) | 0.0363 (12) | |
| C8 | 0.9653 (13) | 0.0361 (6) | 0.5791 (3) | 0.0473 (14) | |
| C9 | 0.0258 (10) | 0.6727 (5) | 0.6633 (2) | 0.0330 (12) | |
| H9A | −0.122640 | 0.615029 | 0.636202 | 0.040* | |
| H9B | 0.160789 | 0.724363 | 0.635096 | 0.040* | |
| C10 | −0.1205 (10) | 0.7763 (5) | 0.7078 (2) | 0.0345 (12) | |
| H10A | −0.249033 | 0.721845 | 0.736309 | 0.041* | |
| H10B | −0.247428 | 0.833281 | 0.682280 | 0.041* | |
| C11 | 0.0872 (10) | 0.8788 (5) | 0.7493 (2) | 0.0379 (13) | |
| H11A | 0.224340 | 0.822607 | 0.772683 | 0.045* | |
| H11B | 0.204592 | 0.939463 | 0.721230 | 0.045* | |
| C12 | −0.0696 (10) | 0.9732 (5) | 0.7969 (3) | 0.0406 (13) | |
| H12A | −0.184891 | 0.912351 | 0.825290 | 0.049* | |
| H12B | −0.208957 | 1.028117 | 0.773504 | 0.049* | |
| C13 | 0.1374 (13) | 1.0785 (6) | 0.8384 (3) | 0.0603 (17) | |
| H13A | 0.275222 | 1.025110 | 0.862082 | 0.090* | |
| H13B | 0.021880 | 1.135181 | 0.868596 | 0.090* | |
| H13C | 0.246599 | 1.141996 | 0.810842 | 0.090* | |
| C14 | 0.7373 (11) | 0.3898 (5) | 0.8135 (2) | 0.0354 (12) | |
| H14A | 0.936232 | 0.418935 | 0.799172 | 0.043* | |
| H14B | 0.716549 | 0.284282 | 0.811463 | 0.043* | |
| C15 | 0.6920 (12) | 0.4506 (5) | 0.8810 (2) | 0.0420 (13) | |
| H15A | 0.828665 | 0.407640 | 0.910064 | 0.050* | |
| H15B | 0.489858 | 0.422903 | 0.893716 | 0.050* | |
| C16 | 0.7393 (14) | 0.6118 (6) | 0.8896 (3) | 0.0523 (16) | |
| H16A | 0.606104 | 0.655330 | 0.859921 | 0.063* | |
| H16B | 0.943233 | 0.639925 | 0.878241 | 0.063* | |
| C17 | 0.6833 (16) | 0.6703 (6) | 0.9585 (3) | 0.0654 (18) | |
| H17A | 0.479985 | 0.640869 | 0.969787 | 0.078* | |
| H17B | 0.817275 | 0.626639 | 0.987940 | 0.078* | |
| C18 | 0.726 (2) | 0.8298 (8) | 0.9686 (3) | 0.111 (3) | |
| H18A | 0.928128 | 0.859964 | 0.958565 | 0.167* | |
| H18B | 0.687568 | 0.859377 | 1.013337 | 0.167* | |
| H18C | 0.590164 | 0.874182 | 0.940628 | 0.167* | |
| H3 | 0.187 (15) | 0.483 (8) | 0.578 (3) | 0.133* | |
| H8 | 1.065 (17) | −0.045 (6) | 0.564 (4) | 0.133* | |
Atomic displacement parameters (Å2) top| | U11 | U22 | U33 | U12 | U13 | U23 |
| O1 | 0.069 (3) | 0.063 (3) | 0.044 (2) | 0.018 (2) | 0.000 (2) | −0.006 (2) |
| N1 | 0.035 (2) | 0.031 (2) | 0.043 (3) | −0.010 (2) | −0.003 (2) | 0.003 (2) |
| C1 | 0.028 (3) | 0.029 (3) | 0.036 (3) | −0.012 (2) | 0.003 (2) | 0.002 (2) |
| O2 | 0.068 (3) | 0.067 (3) | 0.041 (2) | 0.005 (2) | −0.013 (2) | 0.002 (2) |
| C2 | 0.028 (3) | 0.026 (3) | 0.033 (3) | −0.013 (2) | −0.002 (2) | 0.003 (2) |
| O3 | 0.0285 (18) | 0.0289 (18) | 0.040 (2) | −0.0013 (14) | −0.0064 (14) | 0.0051 (15) |
| C3 | 0.024 (2) | 0.026 (3) | 0.038 (3) | −0.011 (2) | −0.004 (2) | 0.006 (2) |
| O4 | 0.0329 (18) | 0.037 (2) | 0.0327 (19) | −0.0043 (15) | −0.0057 (14) | 0.0033 (15) |
| C4 | 0.022 (2) | 0.020 (2) | 0.037 (3) | −0.0078 (19) | 0.002 (2) | 0.005 (2) |
| C5 | 0.023 (2) | 0.027 (3) | 0.036 (3) | −0.011 (2) | −0.002 (2) | 0.005 (2) |
| C6 | 0.025 (3) | 0.027 (3) | 0.040 (3) | −0.007 (2) | −0.004 (2) | 0.006 (2) |
| C7 | 0.037 (3) | 0.029 (3) | 0.041 (3) | −0.010 (2) | 0.001 (2) | 0.001 (2) |
| C8 | 0.061 (4) | 0.039 (3) | 0.041 (3) | −0.003 (3) | 0.002 (3) | 0.001 (3) |
| C9 | 0.023 (2) | 0.032 (3) | 0.044 (3) | 0.003 (2) | −0.003 (2) | 0.007 (2) |
| C10 | 0.025 (3) | 0.029 (3) | 0.050 (3) | −0.003 (2) | −0.002 (2) | 0.012 (2) |
| C11 | 0.030 (3) | 0.030 (3) | 0.054 (3) | −0.002 (2) | 0.000 (2) | 0.007 (2) |
| C12 | 0.032 (3) | 0.030 (3) | 0.060 (3) | −0.001 (2) | −0.002 (2) | 0.009 (2) |
| C13 | 0.053 (4) | 0.045 (4) | 0.080 (4) | 0.001 (3) | −0.010 (3) | −0.010 (3) |
| C14 | 0.037 (3) | 0.032 (3) | 0.037 (3) | −0.005 (2) | −0.007 (2) | 0.008 (2) |
| C15 | 0.048 (3) | 0.042 (3) | 0.035 (3) | −0.009 (2) | −0.005 (2) | 0.007 (2) |
| C16 | 0.069 (4) | 0.042 (3) | 0.044 (3) | −0.014 (3) | −0.007 (3) | 0.001 (3) |
| C17 | 0.096 (5) | 0.052 (4) | 0.047 (4) | 0.002 (4) | −0.008 (3) | −0.005 (3) |
| C18 | 0.210 (11) | 0.063 (5) | 0.057 (5) | 0.001 (6) | −0.002 (5) | −0.013 (4) |
Geometric parameters (Å, º) top
| O1—N1 | 1.215 (5) | C11—C12 | 1.507 (7) |
| N1—O2 | 1.234 (5) | C11—H11A | 0.9900 |
| N1—C2 | 1.469 (6) | C11—H11B | 0.9900 |
| C1—C6 | 1.408 (6) | C12—C13 | 1.542 (7) |
| C1—C2 | 1.418 (7) | C12—H12A | 0.9900 |
| C1—C7 | 1.438 (7) | C12—H12B | 0.9900 |
| C2—C3 | 1.388 (7) | C13—H13A | 0.9800 |
| O3—C4 | 1.359 (5) | C13—H13B | 0.9800 |
| O3—C9 | 1.449 (5) | C13—H13C | 0.9800 |
| C3—C4 | 1.388 (7) | C14—C15 | 1.511 (7) |
| C3—H3 | 0.93 (2) | C14—H14A | 0.9900 |
| O4—C5 | 1.366 (5) | C14—H14B | 0.9900 |
| O4—C14 | 1.440 (6) | C15—C16 | 1.518 (7) |
| C4—C5 | 1.417 (6) | C15—H15A | 0.9900 |
| C5—C6 | 1.379 (6) | C15—H15B | 0.9900 |
| C6—H6 | 0.9500 | C16—C17 | 1.540 (8) |
| C7—C8 | 1.188 (7) | C16—H16A | 0.9900 |
| C8—H8 | 0.95 (2) | C16—H16B | 0.9900 |
| C9—C10 | 1.496 (6) | C17—C18 | 1.503 (9) |
| C9—H9A | 0.9900 | C17—H17A | 0.9900 |
| C9—H9B | 0.9900 | C17—H17B | 0.9900 |
| C10—C11 | 1.529 (6) | C18—H18A | 0.9800 |
| C10—H10A | 0.9900 | C18—H18B | 0.9800 |
| C10—H10B | 0.9900 | C18—H18C | 0.9800 |
| | | |
| O1—N1—O2 | 123.0 (4) | C11—C12—C13 | 113.8 (4) |
| O1—N1—C2 | 120.1 (4) | C11—C12—H12A | 108.8 |
| O2—N1—C2 | 116.9 (4) | C13—C12—H12A | 108.8 |
| C6—C1—C2 | 116.1 (4) | C11—C12—H12B | 108.8 |
| C6—C1—C7 | 117.8 (5) | C13—C12—H12B | 108.8 |
| C2—C1—C7 | 126.1 (4) | H12A—C12—H12B | 107.7 |
| C3—C2—C1 | 122.2 (4) | C12—C13—H13A | 109.5 |
| C3—C2—N1 | 117.1 (4) | C12—C13—H13B | 109.5 |
| C1—C2—N1 | 120.7 (4) | H13A—C13—H13B | 109.5 |
| C4—O3—C9 | 117.2 (4) | C12—C13—H13C | 109.5 |
| C2—C3—C4 | 120.0 (4) | H13A—C13—H13C | 109.5 |
| C2—C3—H3 | 117 (5) | H13B—C13—H13C | 109.5 |
| C4—C3—H3 | 123 (5) | O4—C14—C15 | 107.7 (4) |
| C5—O4—C14 | 117.8 (4) | O4—C14—H14A | 110.2 |
| O3—C4—C3 | 125.0 (4) | C15—C14—H14A | 110.2 |
| O3—C4—C5 | 115.5 (4) | O4—C14—H14B | 110.2 |
| C3—C4—C5 | 119.4 (4) | C15—C14—H14B | 110.2 |
| O4—C5—C6 | 125.7 (4) | H14A—C14—H14B | 108.5 |
| O4—C5—C4 | 114.8 (4) | C14—C15—C16 | 113.9 (4) |
| C6—C5—C4 | 119.5 (4) | C14—C15—H15A | 108.8 |
| C5—C6—C1 | 122.7 (5) | C16—C15—H15A | 108.8 |
| C5—C6—H6 | 118.7 | C14—C15—H15B | 108.8 |
| C1—C6—H6 | 118.7 | C16—C15—H15B | 108.8 |
| C8—C7—C1 | 173.9 (6) | H15A—C15—H15B | 107.7 |
| C7—C8—H8 | 173 (5) | C15—C16—C17 | 112.4 (5) |
| O3—C9—C10 | 107.5 (4) | C15—C16—H16A | 109.1 |
| O3—C9—H9A | 110.2 | C17—C16—H16A | 109.1 |
| C10—C9—H9A | 110.2 | C15—C16—H16B | 109.1 |
| O3—C9—H9B | 110.2 | C17—C16—H16B | 109.1 |
| C10—C9—H9B | 110.2 | H16A—C16—H16B | 107.8 |
| H9A—C9—H9B | 108.5 | C18—C17—C16 | 113.9 (6) |
| C9—C10—C11 | 115.2 (4) | C18—C17—H17A | 108.8 |
| C9—C10—H10A | 108.5 | C16—C17—H17A | 108.8 |
| C11—C10—H10A | 108.5 | C18—C17—H17B | 108.8 |
| C9—C10—H10B | 108.5 | C16—C17—H17B | 108.8 |
| C11—C10—H10B | 108.5 | H17A—C17—H17B | 107.7 |
| H10A—C10—H10B | 107.5 | C17—C18—H18A | 109.5 |
| C12—C11—C10 | 113.3 (4) | C17—C18—H18B | 109.5 |
| C12—C11—H11A | 108.9 | H18A—C18—H18B | 109.5 |
| C10—C11—H11A | 108.9 | C17—C18—H18C | 109.5 |
| C12—C11—H11B | 108.9 | H18A—C18—H18C | 109.5 |
| C10—C11—H11B | 108.9 | H18B—C18—H18C | 109.5 |
| H11A—C11—H11B | 107.7 | | |
| | | |
| C1—C2—N1—O1 | −8.4 (6) | C3—C2—N1—O2 | −8.0 (6) |
Hydrogen-bond geometry (Å, º) top
| D—H···A | D—H | H···A | D···A | D—H···A |
| C3—H3···O2i | 0.93 (2) | 2.88 (3) | 3.793 (7) | 167 (7) |
| C9—H9A···O2i | 0.99 | 2.55 | 3.254 (7) | 128 |
| C8—H8···O1ii | 0.95 (2) | 2.45 (3) | 3.381 (8) | 167 (8) |
| Symmetry codes: (i) −x, −y+1, −z+1; (ii) −x+2, −y, −z+1. |

 Subscribe to Acta Crystallographica Section C: Structural Chemistry
Subscribe to Acta Crystallographica Section C: Structural Chemistry
The full text of this article is available to subscribers to the journal.
If you have already registered and are using a computer listed in your registration details, please email
[email protected] for assistance.
 O,O′ interaction, while the structure of the dibutoxy analogue is dominated by planar ribbons of molecules linked by a similar bifurcated alkoxy sp-C—H
O,O′ interaction, while the structure of the dibutoxy analogue is dominated by planar ribbons of molecules linked by a similar bifurcated alkoxy sp-C—H O,O′ interaction. In contrast, the dipentoxy analogue forms ribbons of molecules alternately connected by a self-complementary sp-C—H
O,O′ interaction. In contrast, the dipentoxy analogue forms ribbons of molecules alternately connected by a self-complementary sp-C—H O2N interaction and a self-complementary sp2-C—H
O2N interaction and a self-complementary sp2-C—H O2N interaction. Disordered solvent was included in the crystals of 1-ethynyl-2-nitro-4,5-dipropoxybenzene and its contribution was removed during refinement.
O2N interaction. Disordered solvent was included in the crystals of 1-ethynyl-2-nitro-4,5-dipropoxybenzene and its contribution was removed during refinement.
 Subscribe to Acta Crystallographica Section C: Structural Chemistry
Subscribe to Acta Crystallographica Section C: Structural Chemistry
 journal menu
journal menu











