issue contents
September 2009 issue
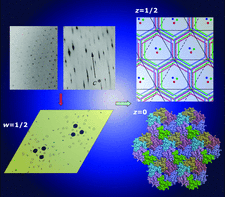
Cover illustration: The diagram illustrates (left) symptoms of lattice translocation disorder (LTD) in diffraction patterns and Patterson maps, and (right) the solution and interpretation of a case of LTD that arose in the determination of a layered hexagonal structure of a carboxysome shell protein (p. 980).
research papers
ACORN2, an improved, more effective version of ACORN, is described and its performance is illustrated by application to a number of known structures, starting with phases from either very small fragments, physical methods or molecular replacement. When the starting fragment is centrosymmetric and gives no definition of an enantiomorph, ACORN2 refinement resolves the problem and converges to a possible solution.
A neutron crystallographic analysis of phosphate-free RNase A has been carried out at 1.7 Å resolution.
PDB reference: ribonuclease A, 3a1r, r3a1rsf
Open  access
access
 access
accessThe crystal structure of the free restriction–modification controller protein C.Esp1396I is reported.
An analysis of the rotational order–disorder structure of fluorescent protein FP480 is presented.
Open  access
access
 access
accessA cyclic olefin homopolymer-based microfluidics system has been established for protein crystallization and in situ X-ray diffraction.
Open  access
access
 access
accessAveraged kick maps are the sum of a series of individual kick maps, where each map is calculated from atomic coordinates modified by random shifts. These maps offer the possibility of an improved and less model-biased map interpretation.
Open  access
access
 access
accessThe first crystal structure of a cereal NTR, a protein involved in seed development and germination, has been determined. The structure is in a conformation that excludes NADPH binding and indicates that a domain reorientation facilitated by Trx binding precedes NADPH binding in the reaction mechanism.
PDB reference: HvNTR2, 2whd, r2whdsf
The crystal structure of the apo decarbamylated form of 2,3-diketo-5-methylthiopentyl-1-phosphate enolase has been solved at 2.3 Å resolution.
Open  access
access
 access
accessA method to accelerate the computation of structure factors from an electron density described by anisotropic and aspherical atomic form factors via fast Fourier transformation is described for the first time.
Corrections for lattice-translocation defects (LTDs) remove the intensity modulation introduced by LTD and decompose the composite structure that is statistically averaged from two LTD-related structures into the original unscrambled structure.
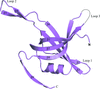
The first crystal structure of a single-stranded DNA-binding protein from a member of the genus Streptomyces has been solved and refined at 2.1 Å resolution. It shows variations in quaternary structure that have previously only been found in mycobacterial SSBs.
PDB reference: single-stranded DNA-binding protein, 3eiv, r3eivsf
A hexagonal layered structure leads to lattice-translocation disorder in crystals of a carboxysome shell protein. Approaches are described for handling translocation problems. In addition, a rare situation is uncovered in which the rotational symmetry in intensity data and in Patterson maps exceeds the rotational symmetry of the crystal itself.
PDB reference: CsoS1C, 3h8y, r3h8ysf
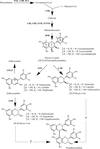
Recombinant grape anthocyanidin reductase synthesizes the naturally rare flavan-3-ol epimers with 2S configuration. Its 2.2 Å resolution crystal structure explains the inactivity of the enzyme in the presence of high salt concentrations.
short communications
An efficient and accurate method has been developed for the preparation of the customized crystallization grid screens employed in protein crystallizations.
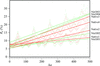
Analysis of a series of diffraction data sets measured from several native as well as nicotinic acid-soaked crystals of trypsin suggests that this potential scavenger does not have any statistically significant effect on the amount of radiation damage incurred in the crystals on X-ray irradiation at 100 K.
letters to the editor
Free 

Comment on Microfibrillar structure of type I collagen in situ by Orgel et al. (2006), Proc. Natl Acad. Sci. USA, 103, 9001–9005.
Free 

A response to the comment on Microfibrillar structure of type I collagen in situ [Okuyama et al. (2009), Acta Cryst. D65, 1007–1008].
book reviews
Free 



 journal menu
journal menu