issue contents
June 2011 issue
Cover illustration: Structure of human R-state aquomethemoglobin at 2.0 Å resolution (Yi et al., p. 647).
structural communications
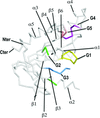
Apo and GDP-bound crystal structures of an essential GTPase, YsxC, from T. maritima were determined to maximal resolutions of 2.3 and 1.9 Å, respectively. Switch I in GDP–YsxC can adopt both an `open' and `closed' conformation, suggesting a mechanism for diffusion of GDP out of the nucleotide-binding pocket.
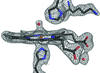
The 2.0 Å resolution crystal structure of tetrameric (αβ)2 R-state human adult aquomethemoglobin reveals axial water coordination at both the α and β heme sites.
PDB reference: R-state aquomethemoglobin, 3p5q
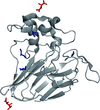
The structure of a recombinant form of the sweet-tasting protein thaumatin I was determined at 1.1 Å resolution and refined to an Rwork of 9.1% and an Rfree of 11.7%. Comparisons with plant thaumatin revealed the electron density of recombinant thaumatin I to be significantly improved, especially around Asn46 and Ser63.
crystallization communications
In this study, the catalytic subunit of E. coli AHAS II was cocrystallized with its cofactors Mg2+, FAD and ThDP using the sitting-drop vapour-diffusion method and the crystals diffracted to 2.80 Å resolution.
The P3 dimerization domain of CheA from E. coli has been crystallized (space group P1, unit-cell parameters a = 59.271, b = 67.674, c = 82.815 Å, α = 77.568, β = 86.073, γ = 64.436°). Diffraction data to 2.80 Å resolution have been collected using synchrotron radiation.
A recombinant form of geraniol dehydrogenase from Backhousia citriodora has been overexpressed in Escherichia coli and purified and crystallized by the sitting-drop method using polyethylene glycol 3350 as a precipitant.
The cloning, overexpression, purification, crystallization and preliminary X-ray crystallographic analysis of phosphoglycerate kinase from S. aureus MRSA252 is reported.
Catechol oxidase from A. oryzae was crystallized by the hanging-drop vapour-diffusion method.
S. Typhimurium CueP has been crystallized. Native data were collected to 2.5 Å resolution.
Factor H-binding protein B from T. denticola was expressed, purified and crystallized. A native data set was collected to 1.8 Å resolution.
Crystals of argininosuccinate lyase from S. mutans were obtained and X-ray data were collected to 2.5 Å resolution in space group R3.
Crystals of dihydrouridine synthase from Thermus thermophilus and its complex with tRNA were obtained and X-ray diffraction data were collected to 1.70 and 3.51 Å resolution, respectively.
Recombinant glucuronic acid dehydrogenase from the halophilic bacterium Chromohalobacter salexigens has been crystallized and X-ray diffraction data collected to a maximum resolution of 2.1 Å.
The Cu-containing nitrite reductase from G. kaustophilus has been overexpressed, purified and crystallized in space group R3. The crystals diffracted to 1.3 Å resolution.
The PH domain of human kindlin-2 was expressed, purified and crystallized. A complete X-ray diffraction data set was obtained at 2.8 Å resolution.
A galactose specific lectin was purified from the seeds of a tropical plant, Spatholobus parviflorus. Its X-ray crystallographic structure was solved by the molecular replacement method.
Recombinant BlaKr has been cocrystallized with kanamycin. A complete data set has been collected to 1.67 Å resolution using synchrotron radiation.
In this study, H. pylori arginase was purified and crystallized in complex with Mn2+ and a diffraction data set was collected to 2.2 Å resolution.
The crystallization and preliminary X-ray crystallographic analysis of the glpX-encoded class II fructose-1,6-bisphosphatase from M. tuberculosis in the apo form is reported.
The purification, characterization and crystallization of a trypsin inhibitor protein isolated from chickpea seeds are reported.
The L. monocytogenes Ca2+-ATPase LMCA1 was crystallized in a Ca2+-free E2–AlF4− form. A complete data set extending to 4.3 Å resolution was collected and a molecular-replacement solution was obtained using sarcoplasmic reticulum Ca2+-ATPase as a search model.
The cloning, expression, purification, crystallization and preliminary X-ray analysis of a ferric binding protein encoded by T. thermophilus HB8 in apo and iron-bound holo states are presented. Four different crystal forms were obtained.
An O-methyltransferase from the ubiquinone-biosynthesis pathway in Escherichia coli, UbiG, with an N-terminal hexahistidine tag has been expressed and crystallized. Crystals grown by the hanging-drop vapour-diffusion method diffracted to 2.00 Å resolution.
The eukaryotic translation initiation factor eIF5BΔN and the eIF5BΔN–eIF1AΔN complex from S. cerevisiae were crystallized. The crystals diffracted to maximum resolutions of 2.45 and 3.3 Å, respectively.


 journal menu
journal menu


























![[publBio]](/logos/publbio.gif)





