issue contents
CCP-FEL: a collection of computer programs for FEL research (August 2016)
Guest editors: Filipe R. N. C. Maia, Thomas A. White, N. Duane Loh and Janos Hajdu
This virtual special issue of Journal of Applied Crystallography brings together a series of specially commissioned articles describing software for free-electron laser research. These articles were published in the journal between April and August 2016.
Cover illustration: CCP-FEL: a collection of computer programs for free-electron laser research.
The latest virtual special issue of Journal of Applied Crystallography presents tools for a range of topics in free-electron laser research such as simulation of experiments, online monitoring of data collection, selection of hits, diagnostics of data quality, data management, data analysis and structure determination for both nanocrystallography and single-particle diffractive imaging.
The WavePropaGator (WPG) package is a new interactive cross-platform open-source software framework for modeling of coherent and partially coherent X-ray wavefront propagation. The WPG addresses the needs of beamline scientists and user groups to facilitate the design, optimization and improvement of X-ray optics to meet their experimental requirements. The paper presents a general description of the package and gives some recent application examples.
Hard X-ray induced dynamics of matter can be simulated with the computational tools XMDYN and XATOM.
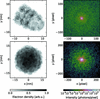
Condor, an open-source simulation tool to predict X-ray scattering amplitudes for flash X-ray imaging experiments, is introduced.
Hummingbird is an open-source scalable Python-based software tool for real-time analysis of diffraction data with the purpose of giving users immediate feedback during their experiments.
This article describes the software package OnDA: online data analysis and feedback for serial X-ray imaging.
An overview of how the well established CFEL–ASG Software Suite (CASS) can be used for serial femtosecond crystallography data is given.
A data processing pipeline for SACLA was developed, based on Cheetah and CrystFEL. Real-time analysis and rapid structure solution were enabled.
The software used to analyze data produced at the Linac Coherent Light Source X-ray free-electron laser is described.
The data acquisition and data management systems used at the Linac Coherent Light Source X-ray free-electron laser are described.
A description is given of a single-particle X-ray imaging reconstruction and simulation package using the expand–maximize–compress algorithm, named Dragonfly.
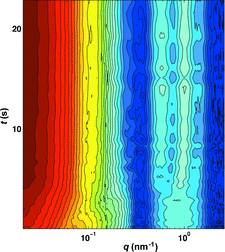
This article describes software for fast model reconstruction from small-angle scattering data, providing real-time feedback on experiments.
Here cppxfel, a software package for integration and post-refinement of serial femtosecond crystallography data, is released.
T. A. White,
V. Mariani,
W. Brehm,
O. Yefanov,
A. Barty,
K. R. Beyerlein,
F. Chervinskii,
L. Galli,
C. Gati,
T. Nakane,
A. Tolstikova,
K. Yamashita,
C. H. Yoon,
K. Diederichs and
H. N. Chapman
Developments in the CrystFEL software suite, for processing diffraction data from `serial crystallography' experiments, are described.
In serial crystallography, CC1/2 may be used as an optimization target, and outlier datasets can be identified on the basis of their influence on the average CC1/2 of the merged data. This leads to the ΔCC1/2 method presented here.
A procedure is presented to model the truncated low pixel counts in micro-electron diffraction (MicroED) images. The correction could extend to any conventional macromolecular X-ray crystallography or X-ray free-electron laser measurements.
An integration optimization, triage and analysis tool (IOTA) is presented, which uses a grid-search approach to maximize the success of indexing and integrating serial X-ray free-electron laser diffraction images. IOTA also includes several useful tools for on-site diffraction data processing.
Related content

Organizers: Filipe R. N. C. Maia and Janos Hajdu
X-ray lasers scheduled to come online in the next few years promise to create a flood of new structural biology data that could overwhelm current processing and analysis methods; this collection describes a series of X-ray free-electron laser datasets generated at the Linac Coherent Light Source, which the organizers hope will help researchers develop new tools and methods to meet these challenges.
Maia, F. R. N. C. & Hajdu, J. (2016). The trickle before the torrent – diffraction data from X-ray lasers. Sci. Data, 3, 160059, doi:10.1038/sdata.2016.59.
Ekeberg, T. et al. (2016). Single-shot diffraction data from the Mimivirus particle using an X-ray free-electron laser. Sci. Data, 3, 160060, doi:10.1038/sdata.2016.60.
Hantke, M. F. et al. (2016). A dataset from flash X-ray imaging of carboxysomes. Sci.
Data, 3, 160061 doi:10.1038/sdata.2016.61.
Munke, A. et al. (2016). Coherent diffraction of single Rice Dwarf Virus particles using hard X-rays at the Linac Coherent Light Source. Sci. Data, 3, 160064, doi:10.1038/sdata.2016.64.
van der Schot, G. et al. (2016). Open data set of live cyanobacterial cells imaged using an X-ray laser. Sci. Data, 3, 160058, doi:10.1038/sdata.2016.58.
White, T. A. et al. (2016). Serial femtosecond crystallography datasets from G protein-coupled receptors. Sci. Data, 3, 160057, 10.1038/sdata.2016.57.
Zhou, X. E. et al. (2016). X-ray laser diffraction for structure determination of the rhodopsin-arrestin complex. Sci. Data, 3, 160021, doi:10.1038/sdata.2016.21.


 access
access access
access access
access access
access access
access access
access access
access access
access access
access access
access access
access access
access access
access


 journal menu
journal menu