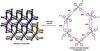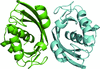issue contents
November 2019 issue

editorial
GENERAL
This Editorial considers the impact of recent work published in IUCrJ and other IUCr journals, as well as the relationship between IUCrJ and the other journals, in terms of where the most cited recent papers are used.
scientific commentaries
CRYO | EM
Nydenova et al. [(2019), IUCrJ, 6, 1086–1098] determine structures of frozen-hydrated protein and nucleic acid assemblies using 100 keV electrons, and describe characteristics of electron microscopes designed to exploit advantages of a lower operating voltage for single-particle cryo-EM.
research letters
MATERIALS | COMPUTATION
In this study, an experimentally synthesized DHM LaNiO3 is discovered with many Dirac cones and complete spin polarization near the Fermi level via first principles; it is also shown that the crystal structures of these materials are strongly correlated with their physical properties.
BIOLOGY | MEDICINE
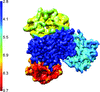
The ∼100 mutations used during the structural determination of GLP-1R suggest multiple mutagenesis strategies, which may find applications in the crystallization of G-protein-coupled receptors and other membrane proteins, and potentially in functional and pharmacological studies by locking proteins into specific conformations.
CRYO | EM
The superior image quality given by the DE64 direct electron detector in counting mode is more important for high-resolution cryo-EM reconstructions than its superior throughput in integrating mode.
research papers
BIOLOGY | MEDICINE
The development and characterization of a surrogate system to support drug discovery targeting a complex heteromeric human glycine receptor is described.
PDB references: acetylcholine-binding protein, wild type, strychnine complex, 5o8t; wild type, nicotine complex, 5o87; variant I, strychnine complex, 5oa0; variant II, HEPES complex, 5oad; variant II, tropisetron complex, 5oaj; variant II, strychnine complex, 5oal; variant III, glycine complex, 5oan; variant III, strychnine complex, 5obg
CRYO | EM
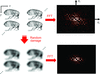
Time-resolved structural studies are becoming an important tool in understanding biological function. Here we describe a cryo-EM grid freezing device capable of rapidly mixing and plunge freezing grids within 10 ms.
CHEMISTRY | CRYSTENG
Download citation


Download citation


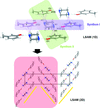
A hydrogen-bond-directed [2+2] photodimerization performed in the solid state is used to generate a head-to-head (HH) cyclobutane photoproduct functionalized with cis-phenolic and cis-pyridyl groups. The photoproduct self-assembles as a pure form in the solid state to form a rare three-dimensional hydrogen-bonded network with a topology that conforms to a mok net. The construction of the HH cyclobutane is achieved using a newly introduced supramolecular protecting-group strategy.
BIOLOGY | MEDICINE
Data from several radiation-damage studies using protein crystals are reanalyzed to determine the local dose- and resolution-dependent decay of diffracted intensity at T = 100 K. The results are inconsistent with long-accepted models and with a proposed linear scaling between maximum tolerable dose and resolution, but are reproduced by a simple, physics-based model.
CRYO | EM
A method (DeepRes) is presented to estimate a new local quality measure for 3D cryoEM maps that adopts the form of a `local resolution' type of information. DeepRes is fully automatic and parameter-free and avoids the issues of most current methods, such as their insensitivity to enhancements owing to B-factor sharpening, among others.
CHEMISTRY | CRYSTENG
Download citation


Download citation


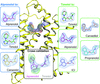
The consistency and variability as well as self-assembly evolution mechanism of cresol–piperazine cocrystals are investigated by experiments and molecular simulation.
BIOLOGY | MEDICINE
The ability to rapidly obtain structures of protein–ligand complexes using X-ray crystallography is central to drug discovery, but the typical cryocooling of samples and the effects of the X-ray beam may distort the observed ligand binding. X-ray free-electron lasers (XFELs) have the promise to solve these issues, but methods to rapidly produce structures of protein–ligand complexes at XFELs have not yet been realized. Here, an efficient solution using high-throughput, fixed-target serial femtosecond crystallography at an XFEL is demonstrated.
CRYO | EM
Several structures were determined by single-particle cryoEM at 100 keV and the implications for future electron cryomicroscopes are considered.
CRYO | EM
Radiation damage is the most fundamental limitation for achieving high resolution in cryo-EM, and is expected to be reduced at liquid-helium temperature. Surprisingly, cryo-EM reconstructions of apoferritin samples cooled with liquid helium showed no improvement in resolution over liquid-nitrogen-cooled samples, but showed substantially more beam-induced particle motion.
BIOLOGY | MEDICINE
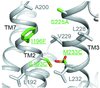
A method is presented for efficient co-crystal structure determination for G protein-coupled receptors taking advantage of serial femtosecond crystallography.
PDB references: β2-adrenergic receptor, complex with alprenolol, 6prz; 6ps2; complex with carazolol, 6ps0; complex with timolol, 6ps1; 6ps6; complex with carvedilol, 6ps3; complex with ICI-118,551, 6ps4; complex with propranolol, 6ps5; adenosine A2A receptor, complex with ZM241385, 6ps7; melatonin receptor MT1, complex with 2-phenylmelatonin, 6ps8
BIOLOGY | MEDICINE
The crystal structure of IdmH from the biosynthetic gene cluster for indanomycin is presented. NMR data show that this enzyme binds its postulated product, indanomycin, and QM/MM modelling of the reaction strongly supports the view that IdmH catalyses indane-ring formation in indanomycin biosynthesis via a Diels–Alder reaction.



 journal menu
journal menu




 access
access