issue contents
September 2023 issue

editorial
GENERAL
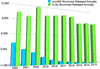
The International Union of Crystallography (IUCr) and its journals are celebrating their 75th anniversary. In this editorial, some of the publication achievements are highlighted and prospects for the future reviewed in an emerging open-access world.
letters to the editor
CRYO | EM
The ongoing debate surrounding the acronym `cryo-EM' is considered.
lead articles
BIOLOGY | MEDICINE
This article reviews phase evaluation in macromolecular crystallography. It accompanies the 2023 Ewald Prize lecture at the 26th IUCr Congress, paying tribute to Paul Ewald's far-reaching influence.
topical reviews
BIOLOGY | MEDICINE
Formate dehydrogenases (FDHs) catalyze the two-electron oxidation of formate to carbon dioxide. This article presents recent progress in the structural analysis of FDHs together with their potential biotechnological applications.
scientific commentaries
BIOLOGY | MEDICINE
A commentary on Wayne Hendrickson's article `Facing the phase problem'.
research papers
CHEMISTRY | CRYSTENG
The competition between contacts in crystal structures is statistically analyzed in several families of molecules using electrostatic energy and enrichment ratio descriptors.
PHYSICS | FELS
This paper presents a neural network model called SpeckleNN for real-time classification of X-ray single-particle imaging (SPI) speckle patterns, ideal for use in high-data-rate facilities like the European XFEL and LCLS-II-HE. It possesses the capability to carry out few-shot classification on novel samples and demonstrates robust performance even in the presence of substantial missing detector area, rendering it an excellent candidate for real-time high-throughput SPI experiments.
CRYO | EM
An update to the MRC image-processing software package is described. It includes improvements to the Ximdisp display program for use on images of helical assemblies or tilt-pair single-particle cryoEM images.
CHEMISTRY | CRYSTENG
Download citation


Download citation


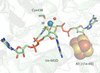
A binding approach based on electronic force density fields was used to describe the structures of the molecule, dimer and crystal of a uracil derivative. Transferable multipole pseudo-atoms were successfully used to reconstruct the inner-crystal electronic force density fields.
CCDC reference: 2259862
BIOLOGY | MEDICINE
The use of intensity based cross-correlation coefficients between half datasets is compared for near atomic resolution protein structure determination and X-ray imaging of cells and tissues.
MATERIALS | COMPUTATION
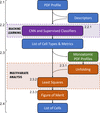
A method to estimate crystal cell parameters directly from a pair distribution function profile, which makes use of machine-learning approaches combined with multivariate analysis and vector superposition techniques, is presented.
BIOLOGY | MEDICINE
The high-resolution crystal structure of AcrIC9 is elucidated and its inhibitory mechanism on the type IC CRISPR–Cas system is unveiled. Analysis and comparison of its structure with structural homologs indicate that AcrIC9 belongs to DNA-mimic Acrs (anti-CRISPR proteins) that directly bind to the cascade complex and hinder the target DNA from binding to the cascade.
PDB reference: AcrIC9, 8hjj


 journal menu
journal menu




 access
access